
| Sequence ID | dm3.chrX |
|---|---|
| Location | 18,372,776 – 18,372,869 |
| Length | 93 |
| Max. P | 0.787178 |

| Location | 18,372,776 – 18,372,869 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 78.12 |
| Shannon entropy | 0.43949 |
| G+C content | 0.51296 |
| Mean single sequence MFE | -28.62 |
| Consensus MFE | -16.04 |
| Energy contribution | -16.48 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.787178 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 18372776 93 + 22422827 CCGGGAAUGCUAAAGGUAAUUCCGAAAUCCCGGCGAGAAUGCCAGGACAUGGCUCAUUUGUCAAGUUGGCUUAGCCAGACUUAGCCGAUUUGC .(((((.(((.....))).))))).......((((((...((((.....))))...))))))(((((((((.((.....)).))))))))).. ( -30.20, z-score = -1.63, R) >droSim1.chrX 14164866 93 + 17042790 CCGAGAAUGCUAAAGGUAAUUCCGAAAUCCCGGCGAGAAUGCCAGGACUUGGCUCAUUUGUCAAGUUGGCUUAGCCAGACUUAGCCGAUUUGC (((((.........((.....))....(((.(((......))).)))))))).........((((((((((.((.....)).)))))))))). ( -27.40, z-score = -1.25, R) >droSec1.super_8 678697 93 + 3762037 CCGAGAAUGCUAAAGGUAAUUCCGAAAUCCCGGCGAGAAUGCCAGGACUUGGCUCAUUUGUCAAGUUGGCUUAGCCAGACUUAGCCGAUUUGC (((((.........((.....))....(((.(((......))).)))))))).........((((((((((.((.....)).)))))))))). ( -27.40, z-score = -1.25, R) >droYak2.chrX 17003747 81 + 21770863 ------------AAGGUAAUUCCGAAAUCCCGGCCAGAAUGCCAGAACUUGGCUCAUUUGUCAAGUUGGCUUAGCCGGACUUAGCCGAUUUAC ------------...((((.((.(.....(((((...((.((((..(((((((......))))))))))))).)))))......).)).)))) ( -25.40, z-score = -2.25, R) >droEre2.scaffold_4690 8690208 92 + 18748788 CCGAGUCUAAUGAAGGUAAUUCCGAAAUCCCGGCAAGAAUGCC-GGACUGGGCUCAUUUGUCAAGUUGGCUUAGCCAGACUUAGCCGAUUUAC ..(((((((.....((.....))......((((((....))))-))..))))))).......(((((((((.((.....)).))))))))).. ( -31.80, z-score = -2.91, R) >droAna3.scaffold_13248 4052626 80 - 4840945 CGGGAGAUAUUCGAGGUAAUACCCGAGUAUCCG-------GCC-GGGUCUGGCUCAUUUGUCAAGCUGGCUUAG--AGGCGGCGGCGGGC--- .....((((((((.((.....))))))))))(.-------(((-(((..((((......))))..)).(((...--.)))..)))).)..--- ( -29.50, z-score = -0.65, R) >consensus CCGAGAAUGCUAAAGGUAAUUCCGAAAUCCCGGCGAGAAUGCCAGGACUUGGCUCAUUUGUCAAGUUGGCUUAGCCAGACUUAGCCGAUUUGC ..............((.....))........(((......(((..((((((((......)))))))))))...)))................. (-16.04 = -16.48 + 0.45)



| Location | 18,372,776 – 18,372,869 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 78.12 |
| Shannon entropy | 0.43949 |
| G+C content | 0.51296 |
| Mean single sequence MFE | -25.60 |
| Consensus MFE | -15.82 |
| Energy contribution | -17.93 |
| Covariance contribution | 2.11 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.715476 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 18372776 93 - 22422827 GCAAAUCGGCUAAGUCUGGCUAAGCCAACUUGACAAAUGAGCCAUGUCCUGGCAUUCUCGCCGGGAUUUCGGAAUUACCUUUAGCAUUCCCGG ((.....((((..(((((((...))))....))).....))))..((((((((......))))))))................))........ ( -26.70, z-score = -1.14, R) >droSim1.chrX 14164866 93 - 17042790 GCAAAUCGGCUAAGUCUGGCUAAGCCAACUUGACAAAUGAGCCAAGUCCUGGCAUUCUCGCCGGGAUUUCGGAAUUACCUUUAGCAUUCUCGG ........((((((((((((...))))....))).......((((((((((((......)))))))))).))........)))))........ ( -28.60, z-score = -1.97, R) >droSec1.super_8 678697 93 - 3762037 GCAAAUCGGCUAAGUCUGGCUAAGCCAACUUGACAAAUGAGCCAAGUCCUGGCAUUCUCGCCGGGAUUUCGGAAUUACCUUUAGCAUUCUCGG ........((((((((((((...))))....))).......((((((((((((......)))))))))).))........)))))........ ( -28.60, z-score = -1.97, R) >droYak2.chrX 17003747 81 - 21770863 GUAAAUCGGCUAAGUCCGGCUAAGCCAACUUGACAAAUGAGCCAAGUUCUGGCAUUCUGGCCGGGAUUUCGGAAUUACCUU------------ ((((.((.(.....((((((((.(((((((((.(......).)))))..))))....))))))))....).)).))))...------------ ( -25.80, z-score = -1.99, R) >droEre2.scaffold_4690 8690208 92 - 18748788 GUAAAUCGGCUAAGUCUGGCUAAGCCAACUUGACAAAUGAGCCCAGUCC-GGCAUUCUUGCCGGGAUUUCGGAAUUACCUUCAUUAGACUCGG ((((.((((((..(((((((...))))....))).....))))...(((-((((....)))))))......)).))))............... ( -25.20, z-score = -1.06, R) >droAna3.scaffold_13248 4052626 80 + 4840945 ---GCCCGCCGCCGCCU--CUAAGCCAGCUUGACAAAUGAGCCAGACCC-GGC-------CGGAUACUCGGGUAUUACCUCGAAUAUCUCCCG ---.......((((..(--((..((((..........)).)).)))..)-)))-------.(((((.(((((......))))).))))).... ( -18.70, z-score = 0.11, R) >consensus GCAAAUCGGCUAAGUCUGGCUAAGCCAACUUGACAAAUGAGCCAAGUCCUGGCAUUCUCGCCGGGAUUUCGGAAUUACCUUUAGCAUUCUCGG .......((((.((.....)).))))...............((((((((((((......)))))))))).))..................... (-15.82 = -17.93 + 2.11)


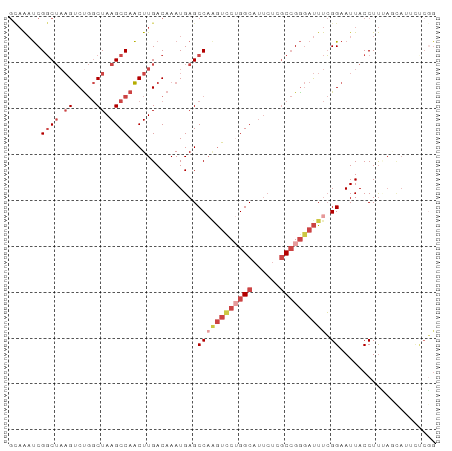
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:56:59 2011