
| Sequence ID | dm3.chrX |
|---|---|
| Location | 18,333,492 – 18,333,604 |
| Length | 112 |
| Max. P | 0.746009 |

| Location | 18,333,492 – 18,333,604 |
|---|---|
| Length | 112 |
| Sequences | 7 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 58.48 |
| Shannon entropy | 0.79592 |
| G+C content | 0.40052 |
| Mean single sequence MFE | -28.07 |
| Consensus MFE | -8.18 |
| Energy contribution | -8.23 |
| Covariance contribution | 0.05 |
| Combinations/Pair | 1.80 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.29 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.746009 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 18333492 112 - 22422827 UCUAGAUGCAACAUCUAGCAUCUGGUUUAGUUAGCAUGCGAGGUUGUUAUUGGCAAUUAGCUUA---GUGGCUUACAGAUCUAAUUAAACCAGAGAUGAAAAAAUCUCAUU-UUUG ...((((((........))))))(((((((((((..((..((((..(((..(((.....)))))---)..)))).))...))))))))))).(((((......)))))...-.... ( -34.90, z-score = -3.71, R) >droSec1.super_8 637122 104 - 3762037 UCUAGAUGACACAUCUAGCAUCUGGUUCAGUUUGCAUGCGAGGUUGUUAUUGGCAAUUA-----------GCUUACAGAUCUAAUUAGGCCAGAGAUUAAAAAAGCUAAUU-AUUG .(((((((...)))))))..((((((..((((....((..(((((((.....)))))..-----------.))..)).....))))..)))))).................-.... ( -22.60, z-score = -0.35, R) >droSim1.chrX 14146807 112 - 17042790 UCUAGAUGACACAUCUAGCAUCUGGUUCAGUUAGCAUGCGAGGUUGUUAUUGGCAAUUAGCUUA---GUGGCUUACAGAUCUAAUUAAGCCAGAGAUUAAAAAAGCUAAUU-AUUG .(((((((...)))))))..(((((((.((((((..((..((((..(((..(((.....)))))---)..)))).))...)))))).))))))).................-.... ( -29.10, z-score = -1.52, R) >droYak2.chrX 16951691 113 - 21770863 UCUAGAUGG---GUCUAACAUCUGGUUUAGUUGGUUUGCGAUGUUGUUAUUGGCAAUUAGCUUAGUUGUAAUCCGCUAAUCUAAUUCAACCAGAGAUUGAAAGCAAUAAUUAAUUG .(((((((.---......)))))))..((((((((((((......(((.((((.((((((.(((((.(....).))))).))))))...)))).))).....)))..))))))))) ( -25.50, z-score = -1.13, R) >droEre2.scaffold_4690 8652443 112 - 18748788 UCUAAAUGGCUGCGAUCUAAUCUGGCUUAGUUGGCAUGCGAUGUUGUUAUUGGCAAUUAGCCUG---CUGGCUUACAGAUCUAAUU-AGCCGGAGAUUGAAAGACCAAAUUAAUAG ......(((((.(((((...(((((((.(((..(((.((...(((((.....)))))..)).))---)..))).............-))))))))))))..)).)))......... ( -33.14, z-score = -1.97, R) >dp4.chrXL_group3a 285085 98 - 2690836 ------------UGCGGACAAAAAAUCCAACUAGC-CGCGAGGUGGGAAUCGGUAUCUC-UUAAUUACUCGGUUGCAUCGCUGAUGCAGCUUCCUCUUCGAAGUAUUUCCUG---- ------------.((((.(..............))-)))((((.((((..(((((....-.....))).))((((((((...)))))))))))).)))).............---- ( -25.54, z-score = -0.85, R) >droPer1.super_39 299245 98 + 745454 ------------UGUGGACAACAAAUCCAACUAGC-CGCGAGGUGGGAAUCGGUAUCUC-UUAAUUACUCGGUUGCAUCGCUGAUGCAGCUUCCUCUUCGAAGUAUUUCCUG---- ------------..((((.......))))....((-..(((((.((((..(((((....-.....))).))((((((((...)))))))))))).)))))..))........---- ( -25.70, z-score = -0.90, R) >consensus UCUAGAUG____AUCUAACAUCUGGUUCAGUUAGCAUGCGAGGUUGUUAUUGGCAAUUAGCUUA___CUGGCUUACAGAUCUAAUUAAGCCAGAGAUUAAAAGAACUAAUU_AUUG ...............(((..(((((((.((((((......(((((((....((.......)).....)))))))......)))))).)))))))..)))................. ( -8.18 = -8.23 + 0.05)


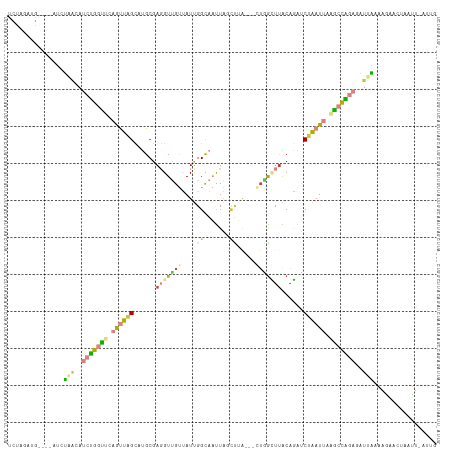
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:56:45 2011