| Sequence ID | dm3.chrX |
|---|---|
| Location | 18,087,678 – 18,087,796 |
| Length | 118 |
| Max. P | 0.738254 |

| Location | 18,087,678 – 18,087,796 |
|---|---|
| Length | 118 |
| Sequences | 8 |
| Columns | 130 |
| Reading direction | forward |
| Mean pairwise identity | 73.49 |
| Shannon entropy | 0.50047 |
| G+C content | 0.42275 |
| Mean single sequence MFE | -31.16 |
| Consensus MFE | -12.69 |
| Energy contribution | -13.16 |
| Covariance contribution | 0.47 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.738254 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 18087678 118 + 22422827 CCAUUGCGCUUCGAUUGCA-GUUUAAGCGCUU-GGCUGUA--UGAGCUUUAAGAGUUU-------UCUUUUAUUGUUUAUU-GCCUCAAGUGCCCCCAUAAGUAGGCAAUACUUUUUGCCGCCUUUCUUU .((((((((((.(((....-))).))))))..-(((...(--(((((..((((((...-------.))))))..)))))).-)))...)))).......(((..(((((......)))))..)))..... ( -32.50, z-score = -2.57, R) >droSim1.chrX_random 189569 118 + 5698898 CCAUUGCGCUUCGAUUGCA-GUUUAAGCGCUU-GGCUAUA--UGAGCUUUAAGAGUUU-------UCUUUUAUUGUUUAUU-GCCUCAAGUGCCCCCAUAAGUAGGCAAUACUUUUUGCCGCCUUUCUUU .((((((((((.(((....-))).))))))..-(((...(--(((((..((((((...-------.))))))..)))))).-)))...)))).......(((..(((((......)))))..)))..... ( -32.50, z-score = -2.77, R) >droSec1.super_8 400340 118 + 3762037 CCAUUGCGCUUCGAUUGCA-GUUUAAGCGCUU-GGCUAUA--UGAGCUUUAAGAGUUU-------UCUUUUAUUGUUUAUU-GCCUCAAGUGCCCCCAUAAGUAGGCAAUACUUUUUGCCGCCUUUCUUU .((((((((((.(((....-))).))))))..-(((...(--(((((..((((((...-------.))))))..)))))).-)))...)))).......(((..(((((......)))))..)))..... ( -32.50, z-score = -2.77, R) >droYak2.chrX 16714926 118 + 21770863 CCAUUGCGCUUCGAUUGCA-GUUUAAGCGCUU-GGCUGUA--UGAGCUUUAAGAGUUU-------UCUUUUAUUGUUUAUU-GCCUCAAGUGCCCCCAUAAGUAGGCAAUACUUUUUGCCGCCAUUCUUU .....((((((.(((....-))).)))))).(-(((.(((--.((((..((((((...-------.))))))..))))(((-(((((..(((....)))..).)))))))......))).))))...... ( -32.60, z-score = -2.57, R) >droEre2.scaffold_4690 8418955 103 + 18748788 CCAUUGCGCUUCGAUUGCA-GUUUAAGCGCCU-GGCUGUA--UGAGCUUUAAGAGUUU-------UCUUUUAUUGUUUAUU-GCCUCAAGUGCCCUCAUAAGUAGGCAAUACUUU--------------- .....((((((.(((....-))).))))))..-(((...(--(((((..((((((...-------.))))))..)))))).-))).....((((..(....)..)))).......--------------- ( -28.80, z-score = -2.14, R) >droAna3.scaffold_13248 3666418 112 - 4840945 --------CUUCGAUUGCA-GUUUAAGCGCCUCGGCUAUU--GGAGCUUUUAAAGUGU-------UCUUUUAUUGUUUAUUUGCCUCGAGUGCGCCCAUAAGUAGGCAAUACUUUUUCCGCCCCCUGUUU --------........(((-(.....((((((((((....--.((((....((((...-------.))))....))))....)))..))).)))).........(((............)))..)))).. ( -24.20, z-score = -0.26, R) >droVir3.scaffold_12970 2789946 118 + 11907090 CCAUCGCAUUUCGAUUGCAUACUGCGACUCGCAGUCUGGUCGUGAGCUUUAAGAGCUAAGGAGUAUUCUUUAUUGUUCAUU-GCACAGAGUGGCCAUAUUAUU-GGCAAUCCUUGUUGCC---------- .....(((.......))).....(((((...((.((((((.((((((...(((((((....)))..))))....)))))).-)).)))).))((((......)-))).......))))).---------- ( -33.70, z-score = -1.31, R) >droMoj3.scaffold_6473 3563953 115 + 16943266 CCAUUGCAUUCGGAUUGCUGGCUUCGAUUCACAGUCGGUUCAUGAGCUUUAAGAGCUUAGGAGUAUUCUUUAUUGUUCAUU-GGCCAGAGUGGCCAUAUUAUU-GGCAAUCCCCGUU------------- ...........((((((((.((((((((.....)))......(((((((...))))))))))))................(-((((.....))))).......-)))))))).....------------- ( -32.50, z-score = -1.24, R) >consensus CCAUUGCGCUUCGAUUGCA_GUUUAAGCGCUU_GGCUGUA__UGAGCUUUAAGAGUUU_______UCUUUUAUUGUUUAUU_GCCUCAAGUGCCCCCAUAAGUAGGCAAUACUUUUUGCCGCC_UUCUUU .............(((((........((((...(((......(((((..((((((...........))))))..)))))...)))....))))............))))).................... (-12.69 = -13.16 + 0.47)


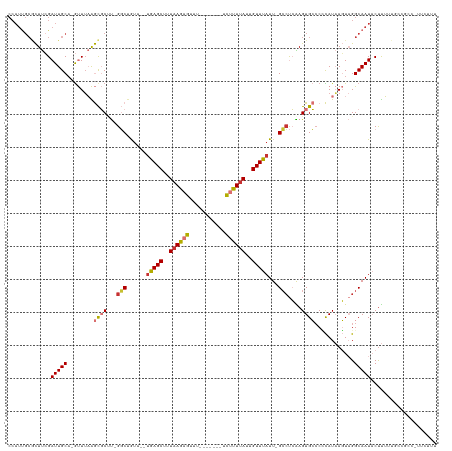
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:55:32 2011