
| Sequence ID | dm3.chrX |
|---|---|
| Location | 17,899,484 – 17,899,581 |
| Length | 97 |
| Max. P | 0.606573 |

| Location | 17,899,484 – 17,899,581 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 76.11 |
| Shannon entropy | 0.44651 |
| G+C content | 0.51924 |
| Mean single sequence MFE | -31.17 |
| Consensus MFE | -18.73 |
| Energy contribution | -19.43 |
| Covariance contribution | 0.70 |
| Combinations/Pair | 1.36 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.606573 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
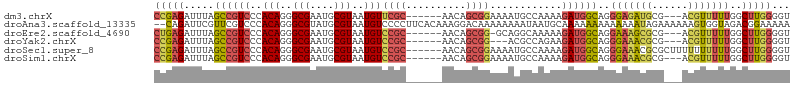
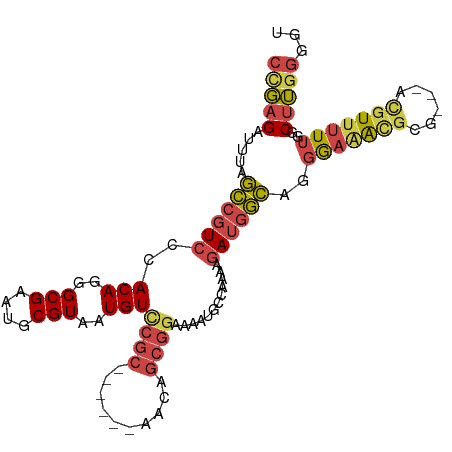
>dm3.chrX 17899484 97 - 22422827 CCGAGAUUUAGCCGUCCCACAGGGCGAAUGCGUAAUGUUCGC------AACAGCGGAAAAUGCCAAAAGAUGGCAGGGAGAUGCG---ACGUUUUUGGCUUGGGGU ...............(((.((((.((((.((((....(((((------....)))))...(((((.....)))))..........---)))).)))).))))))). ( -29.90, z-score = -0.43, R) >droAna3.scaffold_13335 1607064 104 + 3335858 --CAGAUUCGUUCGUCCCACAGGGCGUAUGCGUAAUGUCCCCUUCACAAAGGACAAAAAAAAUAAUGCAAAAAAAAAAAAAUAGAAAAAAGUGGUAGACGGAAAAA --........((((((((((.((((((.......))))))((((....))))......................................))))..)))))).... ( -20.40, z-score = -1.95, R) >droEre2.scaffold_4690 8227295 96 - 18748788 CUGAGAUUUAGCCGUCCCACAGGGCGAAUGCGUAAUGUCCGC------AACAGCGG-GCAGGCAAAAAGAUGGCAGGAAAGCGCG---ACGUUUUUGGCUUGGGGU (..((.....(((((((....))))...(((....(((((((------....))))-))).))).......)))..(((((((..---.)))))))..))..)... ( -31.80, z-score = -0.55, R) >droYak2.chrX 16518980 94 - 21770863 CCGAGAUUUAGCCGUCCCACAGGGCGAAUGCGUAAUGUCCGC------AACAGCGG---ACGCCAGAAGAUGGCAGGGAAACGCG---ACGUUUUUGGCUUGGGGU ...............(((.((((.((((.((((...((((((------....))))---))((((.....))))..(....)...---)))).)))).))))))). ( -37.40, z-score = -2.25, R) >droSec1.super_8 200156 100 - 3762037 CCGAGAUUUAGCCGUCCCACAGGGCGAAUGCGUAAUGUCCGC------AACAGCGGAAAAUGCCAAAAGAUGGCAGGGAAACGCGCUUUUUUUUUUGGCUUGGGGU ((((......(((((((....)))))...))......(((((------....)))))....(((((((((.(((..(....)..)))...)))))))))))))... ( -33.60, z-score = -0.92, R) >droSim1.chrX 13923304 97 - 17042790 CCGAGAUUUAGCCGUCCCACAGGGCGAAUGCGUAAUGUCCGC------AACAGCGGAAAAUGCCAAAAGAUGGCAGGGAAACGCG---ACGUUUUUGGCUUGGGGU ...............(((.((((.((((.((((....(((((------....)))))...(((((.....))))).(....)...---)))).)))).))))))). ( -33.90, z-score = -1.38, R) >consensus CCGAGAUUUAGCCGUCCCACAGGGCGAAUGCGUAAUGUCCGC______AACAGCGGAAAAUGCCAAAAGAUGGCAGGGAAACGCG___ACGUUUUUGGCUUGGGGU (((((.....((((((..(((..(((....)))..)))((((..........))))............))))))..(((((((......)))))))..)))))... (-18.73 = -19.43 + 0.70)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:54:48 2011