
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 11,737,796 – 11,737,909 |
| Length | 113 |
| Max. P | 0.642139 |

| Location | 11,737,796 – 11,737,909 |
|---|---|
| Length | 113 |
| Sequences | 7 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 70.38 |
| Shannon entropy | 0.56022 |
| G+C content | 0.45274 |
| Mean single sequence MFE | -26.73 |
| Consensus MFE | -12.99 |
| Energy contribution | -13.75 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.642139 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
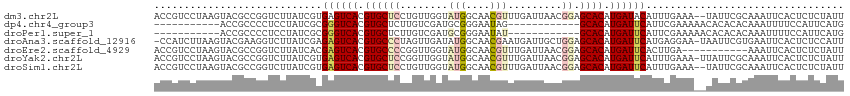
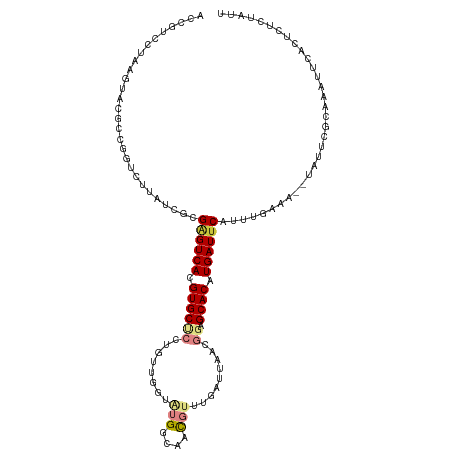
>dm3.chr2L 11737796 113 - 23011544 ACCGUCCUAAGUACGCCGGUCUUAUCGUGAGUCACGUGCUCCUGUUGGUAUGGCAACGUUUGAUUAACGGAGCACAUGAUACAUUUGAAA--UAUUCGCAAAUUCACUCUCUAUU ((((.(........).))))......((((((((.(((((((.(((((((((....)))...))))))))))))).))))..(((((...--......)))))))))........ ( -29.60, z-score = -1.55, R) >dp4.chr4_group3 2608958 92 - 11692001 -----------ACCGCCCCUCCUAUCGCGGGUCACGUGCUCUUGUCGAUGCGGGAAUAG------------GCACAUGAUUCAUUCGAAAAACACACACAAAUUUUCCAUUCAUG -----------.((((..........))))((((.((((((((((....))))))....------------)))).))))................................... ( -17.00, z-score = 0.54, R) >droPer1.super_1 4083178 92 - 10282868 -----------ACCGCCCCUCCUAUCGCGGGUCACGUGCUCUUGUCGAUGCGGGAAUAU------------GCACAUGAUUCAUUCGAAAAACACACACAAAUUUUCCAUUCAUG -----------.((((..........))))((((.((((((((((....))))))....------------)))).))))................................... ( -17.00, z-score = 0.23, R) >droAna3.scaffold_12916 9445928 113 + 16180835 -CCAUCUUAAGUACGAAGGUCUUAUCGAGAGUCACGUGCCCUAGUUGAUAUGGCAACGAAUGAUUGCUGGAGCACAUGAUUCAUGAGGAA-UAAUUCGUGAAUUCACUCUCCAUU -......((((.((....))))))..((((((...((((.((((((.((.((....)).)).)..))))).))))..((((((((((...-...)))))))))).)))))).... ( -34.20, z-score = -2.40, R) >droEre2.scaffold_4929 12962214 104 + 26641161 ACCGUCCUAAGUACGCCGGUCUUAUCACGAGUCACGUGCCCCGGUUGGUAUGGCAACGUUUGAUUAACGGAGCACAUGAUUCACUUGA-----------AAAUUCACUCUCUAUU ...((..((((.((....))))))..))((((((.((((.(((((..(.(((....))))..))...))).)))).))))))......-----------................ ( -25.90, z-score = -0.65, R) >droYak2.chr2L 8157008 114 - 22324452 ACCGUCCUAAGUACGCCGGUCUUAUCGUGAGUCACGUGCUCCGGUUGGUAUGGCAACGUUUGAUUAACGGAGCACAUGAUUCAUUUGAAA-UUAUUCGCAAAUUCACUCUCUAUU ..............((..((..(.((((((((((.((((((((((..(.(((....))))..))...)))))))).))))))))..)).)-..))..))................ ( -31.80, z-score = -2.01, R) >droSim1.chr2L 11547869 113 - 22036055 ACCGUCCUAAGUACGCCGGUCUUAUCGUGAGUCACGUGCUCCUGUUGGUAUGGCAACGUUUGAUUAACGGAGCACAUGAUUCAUUUGAAA--UAUUCGCAAAUUCACUCUCUAUU ((((.(........).))))......((((((((.(((((((.(((((((((....)))...))))))))))))).))))))))......--....................... ( -31.60, z-score = -2.27, R) >consensus ACCGUCCUAAGUACGCCGGUCUUAUCGCGAGUCACGUGCUCCUGUUGGUAUGGCAACGUUUGAUUAACGGAGCACAUGAUUCAUUUGAAA__UAUUCGCAAAUUCACUCUCUAUU ............................((((((.((((.((.......(((....))).........)).)))).))))))................................. (-12.99 = -13.75 + 0.76)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:33:29 2011