
| Sequence ID | dm3.chrX |
|---|---|
| Location | 17,655,116 – 17,655,201 |
| Length | 85 |
| Max. P | 0.820680 |

| Location | 17,655,116 – 17,655,201 |
|---|---|
| Length | 85 |
| Sequences | 11 |
| Columns | 86 |
| Reading direction | forward |
| Mean pairwise identity | 77.57 |
| Shannon entropy | 0.46969 |
| G+C content | 0.54311 |
| Mean single sequence MFE | -27.98 |
| Consensus MFE | -15.71 |
| Energy contribution | -15.31 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.36 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.820680 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 17655116 85 + 22422827 UGCAGCGGCGCAGCGGGGCGUCGGAGGUGUGAAAACCCUGCUGGGUCUUAAAUGCAAGC-UAAUAAGCAGCGGACUGCAAGUCAGG ((((((.(((((..(((((.(((((((.((....))))).)))))))))...)))..((-(....))).)).).)))))....... ( -31.80, z-score = -1.05, R) >droSim1.chrX 13749496 85 + 17042790 UGCAGCGGCGCAGCGGGGCGUCGGAGGUGUGAAAACCCUGCUGGGUCUUAAAUGCAAGC-UAAUAAGCAGCGGACUGCAAGUCAGG ((((((.(((((..(((((.(((((((.((....))))).)))))))))...)))..((-(....))).)).).)))))....... ( -31.80, z-score = -1.05, R) >droSec1.super_17 1501473 85 + 1527944 UGCAGCGGCGCAGCGGGGCGUCGGAGGUGUGAAAACCCUGCUGGGUCUUAAAUGCAAGC-UAAUAAGCAGCGGACUGCAAGUCAGG ((((((.(((((..(((((.(((((((.((....))))).)))))))))...)))..((-(....))).)).).)))))....... ( -31.80, z-score = -1.05, R) >droYak2.chrX 16284322 85 + 21770863 UGCAGCGGCGCAGUGGGGCGUCGGAGGUGUGAAAACCCUGCUGGGUCUUAAAUGCAAGC-UAAUAAGCAGCGGACUGCAAGUCAGG ((((((.(((((.((((((.(((((((.((....))))).))))))))))..)))..((-(....))).)).).)))))....... ( -32.20, z-score = -1.49, R) >droEre2.scaffold_4690 7996886 85 + 18748788 UGCAGCGGCGCAGUGGGGCGUCGGAGGUGUGAAAACCCUGCUGGGUCUUAAAUGCAAGC-UAAUAAGCAGCGGACUGCAAGUCAGG ((((((.(((((.((((((.(((((((.((....))))).))))))))))..)))..((-(....))).)).).)))))....... ( -32.20, z-score = -1.49, R) >droAna3.scaffold_13335 2953110 68 - 3335858 -------UCGCAGUGGGGCGUCGGAGGUGUGAAAACCCUGCUGGGUCUUAAAUGCAAGC-UAAUAAGAAGCACAGG---------- -------..(((.((((((.(((((((.((....))))).))))))))))..)))..((-(.......))).....---------- ( -20.20, z-score = -1.01, R) >dp4.chrXL_group1e 12043510 78 + 12523060 AGCAGAGGCGCAGUGGGGCGUCAGAGGUGUGAAAACCCUGCUGGGUCUUAA-UGCAAGC-UAAUAAGAAGCAGCACCGAA------ .((((((((.(((..(((..(((......)))...)))..))).)))))..-)))..((-(.......))).........------ ( -27.30, z-score = -1.50, R) >droPer1.super_14 601207 71 - 2168203 AGCAGAGGCGCAGUGGGGCGUCAGAGGUGUGAAAACCCUGCUGGGUCUUAA-UGCAAGC-UAAUAAGAAGCAG------------- .((((((((.(((..(((..(((......)))...)))..))).)))))..-)))..((-(.......)))..------------- ( -27.30, z-score = -2.86, R) >droWil1.scaffold_181096 4008129 83 + 12416693 AGACGCAGCCCAGUGGGGCGCAAAAGGUGUGAAAACCCUGCUGGGUUUUAA-UGCAAGCCUAAUAAGAAGCAGUACAAGUCAAG-- .((((((((((((..(((((((.....))))....)))..)))))).....-)))..((..........)).......)))...-- ( -27.50, z-score = -2.03, R) >droGri2.scaffold_14853 6745506 74 + 10151454 ------GACGCAGUGGGGUGCAAAAGGUGUGAAAACCCUGCUGCGUUUUAAAUGCAAGC-UAAUAAGAACGAGAACGA--CAA--- ------(((((((..((((.((.......))...))))..)))))))............-..................--...--- ( -21.60, z-score = -2.30, R) >droVir3.scaffold_12928 5070360 74 - 7717345 ------GACGCAGUGGGGCGCAAAAGGUGUGAAAACCCUGCUGCGUUUUAAAUGCAAGC-UAAUAAGAACAGGCACAG--CCA--- ------(((((((..(((((((.....))))....)))..)))))))............-...........(((...)--)).--- ( -24.10, z-score = -1.37, R) >consensus UGCAGCGGCGCAGUGGGGCGUCGGAGGUGUGAAAACCCUGCUGGGUCUUAAAUGCAAGC_UAAUAAGAAGCGGACUGCAAGUC___ ......(((.((((((((.................)))))))).)))....................................... (-15.71 = -15.31 + -0.40)


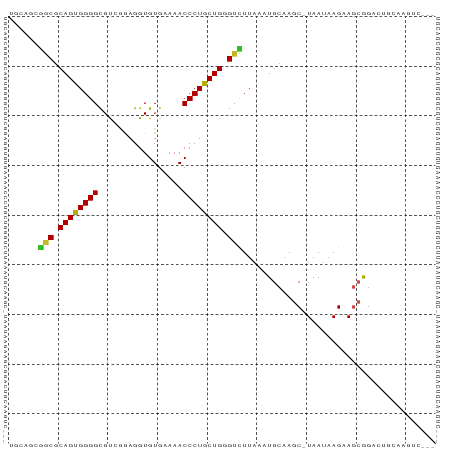
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:54:11 2011