
| Sequence ID | dm3.chrX |
|---|---|
| Location | 17,623,244 – 17,623,361 |
| Length | 117 |
| Max. P | 0.869858 |

| Location | 17,623,244 – 17,623,361 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.82 |
| Shannon entropy | 0.22664 |
| G+C content | 0.38680 |
| Mean single sequence MFE | -26.70 |
| Consensus MFE | -18.21 |
| Energy contribution | -18.13 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.869858 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 17623244 117 + 22422827 --ACUAGUAAUCGUAAUCGCUGGGAUCUGGCAAGCUCUCUUUUAUGAGCACAUUUACAGCUACUAAUUUGCGCUCAAAUUACCAGAAAUUAACACAAUUUCACUU-GCGAUAACAUCAGC --..........((.(((((((((((.((((..((((........)))).........((.........)))).)).))).)))((((((.....))))))....-))))).))...... ( -24.90, z-score = -2.03, R) >droSim1.chrX 13726937 118 + 17042790 ACUCUAGUAAUCGUAAUCGCUGGGAUCUGGCAAGCUCUCUUUUAUGAGCCCAUUUACAGCUACUAAUU-GCGCUCAAAUUACCAGAAAUUAACACAAUUUCCCUU-GCGAUAAUAUCAGC ...............(((((.(((((((((...((((........)))).........((........-))..........))))).............))))..-)))))......... ( -24.61, z-score = -1.52, R) >droSec1.super_17 1468848 118 + 1527944 ACUCUAGCAAUCGUAAUCGCUGGGAUCUGGCAAGCUCUCUUUUAUGAGCCCAUUUACAGCUACUAAUU-GAGCUGAAAUUACCAGAAAUUAACACAAUUUUCUUU-GCGAUAAUAUAAGC ...............(((((..((((((((...((((........)))).......(((((.......-.)))))......)))))((((.....)))).)))..-)))))......... ( -25.70, z-score = -1.49, R) >droYak2.chrX 16251044 111 + 21770863 --------ACAGAUAAUCGCUGGAAUCUGGGAAGCUCUCUUUUAUGAGCUCAUUUGCAGCUACUAAUU-GCGCGCAAAUUACCAGAAAUAAACACAAUUUCCCUUUGCGAUAAUAUCAGC --------...(((((((((.(((((((((((.((((........))))))((((((.((........-))..))))))..)))))............))))....)))))..))))... ( -31.40, z-score = -3.39, R) >droEre2.scaffold_4690 7958278 110 + 18748788 --------ACAAAUAAUCGCUGGGAUCUGGGAAGCUCUCUUUUAUGAGCUCAUUUACAGCUACUAAUU-GCACUCAAAUUACCAGAAAUUAACACAACUUCCCUU-GCGAUAUUAUCAGC --------.......(((((.(((((((((((.((((........)))))).......((........-))..........))))).............))))..-)))))......... ( -26.91, z-score = -3.68, R) >consensus ___CUAG_AAUCGUAAUCGCUGGGAUCUGGCAAGCUCUCUUUUAUGAGCCCAUUUACAGCUACUAAUU_GCGCUCAAAUUACCAGAAAUUAACACAAUUUCCCUU_GCGAUAAUAUCAGC ............((.(((((.(((((((((...((((........))))..............(((((........)))))))))).............))))...))))).))...... (-18.21 = -18.13 + -0.08)

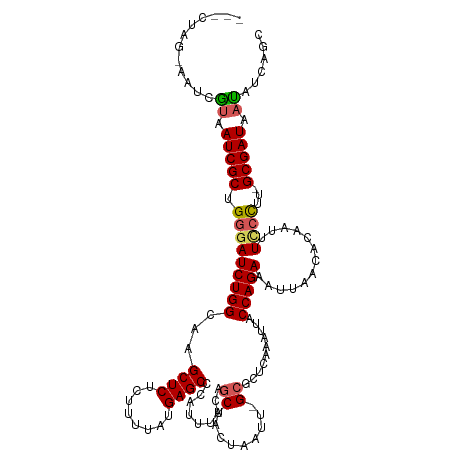

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:54:01 2011