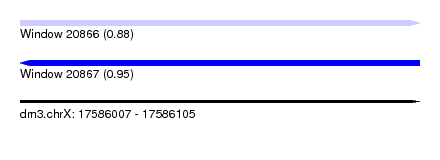

| Sequence ID | dm3.chrX |
|---|---|
| Location | 17,586,007 – 17,586,105 |
| Length | 98 |
| Max. P | 0.950920 |

| Location | 17,586,007 – 17,586,105 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 75.00 |
| Shannon entropy | 0.40591 |
| G+C content | 0.44789 |
| Mean single sequence MFE | -30.30 |
| Consensus MFE | -21.28 |
| Energy contribution | -21.28 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.05 |
| SVM RNA-class probability | 0.881080 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 17586007 98 + 22422827 ---------------UUUUGAAUGCAUUAAACUUGUCUAGCAAUUCGUUCUAGGCUGGAUAACAGGGUCACACUGCUCAGGCUGUGUGACCAAGCGACAAUGAAAAGCAUUGC------- ---------------.......(((.......(((((((((............)))))))))...((((((((.((....)).))))))))..))).(((((.....))))).------- ( -32.00, z-score = -2.69, R) >droEre2.scaffold_4690 7921631 112 + 18748788 AUGCAGACCUCAGUCUUUUAAAUGCACAAAUCUUGUCUUGACUUUCGUUCUAGG-UGGAUAGCAGGGUCACACUACUCAUGCUGUGUGACCAAGCAUCAAUGAAAAGCAUUGC------- ..((((((....))))....(((((........(((((...((........)).-.)))))((..((((((((..(....)..))))))))..))...........)))))))------- ( -28.90, z-score = -0.66, R) >droYak2.chrX 16213949 97 + 21770863 -----------------------AUAGCAACUUUAGUCAUGGGCUUCUUCUAGGCUGGGCAGCAGGGUCACACUGCACAUGCUGUGUGACCAAGCUUCAAUGAAUAGCAUUGCUAAAGAU -----------------------.((((((..(((.((((.(((((.......(((....)))..((((((((.((....)).)))))))))))))...)))).)))..))))))..... ( -35.00, z-score = -2.34, R) >droSec1.super_17 1431963 98 + 1527944 ---------------CUUUGAAUACAUAAAACUUGUCUUGCAAUUCGUUCUAGGCUGGGUAGCAGGGUCACACUGCUCAGGCUGUGUGACCAAGCGUCAAUGAAAAGCAUUGC------- ---------------...............((((((((.((.....))...)))).)))).((..((((((((.((....)).))))))))..))..(((((.....))))).------- ( -27.80, z-score = -0.97, R) >droSim1.chrX_random 4522158 98 + 5698898 ---------------UUUUGAAUACAUAAAACUUGUCUUGCAAUUCGUUCUAGGCUGGGUAGCAGGGUCACACUGCUCAGGCUGUGUGACCAAGCGUCAAUGAAAAGCAUUGC------- ---------------...............((((((((.((.....))...)))).)))).((..((((((((.((....)).))))))))..))..(((((.....))))).------- ( -27.80, z-score = -1.05, R) >consensus _______________UUUUGAAUACAUAAAACUUGUCUUGCAAUUCGUUCUAGGCUGGGUAGCAGGGUCACACUGCUCAGGCUGUGUGACCAAGCGUCAAUGAAAAGCAUUGC_______ ..........................................(((((........))))).((..((((((((.((....)).))))))))..))..(((((.....)))))........ (-21.28 = -21.28 + -0.00)



| Location | 17,586,007 – 17,586,105 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 75.00 |
| Shannon entropy | 0.40591 |
| G+C content | 0.44789 |
| Mean single sequence MFE | -27.90 |
| Consensus MFE | -20.72 |
| Energy contribution | -20.40 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.57 |
| SVM RNA-class probability | 0.950920 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 17586007 98 - 22422827 -------GCAAUGCUUUUCAUUGUCGCUUGGUCACACAGCCUGAGCAGUGUGACCCUGUUAUCCAGCCUAGAACGAAUUGCUAGACAAGUUUAAUGCAUUCAAAA--------------- -------(.(((((......(((((((..((((((((.((....)).))))))))..)).....(((............))).))))).......))))))....--------------- ( -26.72, z-score = -1.79, R) >droEre2.scaffold_4690 7921631 112 - 18748788 -------GCAAUGCUUUUCAUUGAUGCUUGGUCACACAGCAUGAGUAGUGUGACCCUGCUAUCCA-CCUAGAACGAAAGUCAAGACAAGAUUUGUGCAUUUAAAAGACUGAGGUCUGCAU -------(((((((...((...(((((..((((((((.((....)).))))))))..)).)))..-.((.((.(....))).))....)).....)))))....((((....)))))).. ( -32.20, z-score = -1.70, R) >droYak2.chrX 16213949 97 - 21770863 AUCUUUAGCAAUGCUAUUCAUUGAAGCUUGGUCACACAGCAUGUGCAGUGUGACCCUGCUGCCCAGCCUAGAAGAAGCCCAUGACUAAAGUUGCUAU----------------------- .....(((((((..((.((((....((((((((((((.((....)).))))))))..(((....))).......))))..)))).))..))))))).----------------------- ( -31.50, z-score = -2.57, R) >droSec1.super_17 1431963 98 - 1527944 -------GCAAUGCUUUUCAUUGACGCUUGGUCACACAGCCUGAGCAGUGUGACCCUGCUACCCAGCCUAGAACGAAUUGCAAGACAAGUUUUAUGUAUUCAAAG--------------- -------.(((((.....)))))..((..((((((((.((....)).))))))))..))......((.((((((..............)))))).))........--------------- ( -24.54, z-score = -1.25, R) >droSim1.chrX_random 4522158 98 - 5698898 -------GCAAUGCUUUUCAUUGACGCUUGGUCACACAGCCUGAGCAGUGUGACCCUGCUACCCAGCCUAGAACGAAUUGCAAGACAAGUUUUAUGUAUUCAAAA--------------- -------.(((((.....)))))..((..((((((((.((....)).))))))))..))......((.((((((..............)))))).))........--------------- ( -24.54, z-score = -1.29, R) >consensus _______GCAAUGCUUUUCAUUGACGCUUGGUCACACAGCCUGAGCAGUGUGACCCUGCUACCCAGCCUAGAACGAAUUGCAAGACAAGUUUUAUGUAUUCAAAA_______________ ........(((((.....)))))..((..((((((((.((....)).))))))))..))............................................................. (-20.72 = -20.40 + -0.32)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:53:49 2011