
| Sequence ID | dm3.chrX |
|---|---|
| Location | 17,491,002 – 17,491,111 |
| Length | 109 |
| Max. P | 0.779859 |

| Location | 17,491,002 – 17,491,111 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 76.06 |
| Shannon entropy | 0.43607 |
| G+C content | 0.60707 |
| Mean single sequence MFE | -41.22 |
| Consensus MFE | -26.52 |
| Energy contribution | -28.52 |
| Covariance contribution | 2.00 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.23 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.664256 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 17491002 109 + 22422827 CAUGGACCUGACCCACCGACCCACCACCACACGGCCGGUGGUAACCCCCAGUUAUGGUGGUGGCCCCCAAUUGGCCCAGGUCCAUGAUGUUGAACAGGCAGUGGAGCUG ((((((((((..((((((..(((((.((....))..)))))((((.....)))))))))).((((.......))))))))))))))..........(((......))). ( -45.00, z-score = -1.28, R) >droSim1.chrX 3580455 109 - 17042790 CAUGGACCUGACCCACCGACCCACCACCACCCGGCCGGUGGUGACCCCCAGUUAUGGUGGUAGCCCCGAAUUGGCGCAGGUCCAUGAUGUUGAACAGGCAGUGGAGCUG ((((((((((..((((((...(((((((........)))))))((.....))..))))))..(((.......))).))))))))))..........(((......))). ( -43.40, z-score = -1.39, R) >droSec1.super_17 1333839 109 + 1527944 CAUGGACCUGACCCACCGACCCACCACCACCCGGCCGGUGGUGACCCCCAGUUAUGGUGGUAGCCCCGAAUUGGCGCAGGUCCAUGAUGUUGAACAGGCAGUGGAGCUG ((((((((((..((((((...(((((((........)))))))((.....))..))))))..(((.......))).))))))))))..........(((......))). ( -43.40, z-score = -1.39, R) >droYak2.chrX 16118677 109 + 21770863 CAUGGACCUAACCCACCGACCCACCACCAGCCGGCCGGUGGUGACCCCCAGUUAUGGUGGUGGCCCCGAACUGGCGCAGGUCGCUGAUGUUGACCAGGCGGUGGAGCUG ..(((.......)))..(..(((((.((.(((((.(((.(((.(((.(((....))).))).))))))..)))))...((((((....).))))).)).)))))..).. ( -48.30, z-score = -1.23, R) >droWil1.scaffold_180702 45686 103 - 4511350 CAUGGAUUUGACUUUAAGGCCAACAACGG--CAGCCAGU--CGAUCUUCAAUGGUCACAACGUCGAUGGGAGAG-ACACUUUCA-GAUGUUGAUCACAAAGACGAGGAG .......((((((.....(((......))--)....)))--)))(((((........(((((((..((..((..-...))..))-))))))).((.....)).))))). ( -26.00, z-score = -0.84, R) >consensus CAUGGACCUGACCCACCGACCCACCACCACCCGGCCGGUGGUGACCCCCAGUUAUGGUGGUGGCCCCGAAUUGGCGCAGGUCCAUGAUGUUGAACAGGCAGUGGAGCUG ((((((((((..(((......(((((((........)))))))(((.(((....))).)))..........)))..))))))))))..........(((......))). (-26.52 = -28.52 + 2.00)


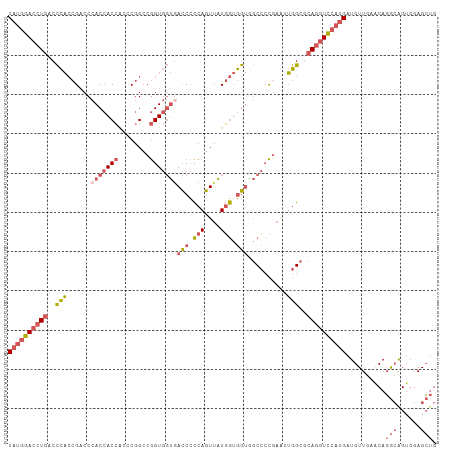
| Location | 17,491,002 – 17,491,111 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 76.06 |
| Shannon entropy | 0.43607 |
| G+C content | 0.60707 |
| Mean single sequence MFE | -45.40 |
| Consensus MFE | -32.00 |
| Energy contribution | -34.08 |
| Covariance contribution | 2.08 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.67 |
| SVM RNA-class probability | 0.779859 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 17491002 109 - 22422827 CAGCUCCACUGCCUGUUCAACAUCAUGGACCUGGGCCAAUUGGGGGCCACCACCAUAACUGGGGGUUACCACCGGCCGUGUGGUGGUGGGUCGGUGGGUCAGGUCCAUG ..((......))...........(((((((((((.(((.....((.(((((((((((.((((((....)).))))...))))))))))).))..))).))))))))))) ( -56.00, z-score = -2.38, R) >droSim1.chrX 3580455 109 + 17042790 CAGCUCCACUGCCUGUUCAACAUCAUGGACCUGCGCCAAUUCGGGGCUACCACCAUAACUGGGGGUCACCACCGGCCGGGUGGUGGUGGGUCGGUGGGUCAGGUCCAUG ..((......))...........((((((((((.(((....(.((.((((((((((..((((((....)).))))....)))))))))).)).)..))))))))))))) ( -51.50, z-score = -1.85, R) >droSec1.super_17 1333839 109 - 1527944 CAGCUCCACUGCCUGUUCAACAUCAUGGACCUGCGCCAAUUCGGGGCUACCACCAUAACUGGGGGUCACCACCGGCCGGGUGGUGGUGGGUCGGUGGGUCAGGUCCAUG ..((......))...........((((((((((.(((....(.((.((((((((((..((((((....)).))))....)))))))))).)).)..))))))))))))) ( -51.50, z-score = -1.85, R) >droYak2.chrX 16118677 109 - 21770863 CAGCUCCACCGCCUGGUCAACAUCAGCGACCUGCGCCAGUUCGGGGCCACCACCAUAACUGGGGGUCACCACCGGCCGGCUGGUGGUGGGUCGGUGGGUUAGGUCCAUG .(((.(((((((.((((....))))))((((..(.((((((..((.......))..)))))).)..(((((((((....)))))))))))))))))))))......... ( -49.90, z-score = -0.72, R) >droWil1.scaffold_180702 45686 103 + 4511350 CUCCUCGUCUUUGUGAUCAACAUC-UGAAAGUGU-CUCUCCCAUCGACGUUGUGACCAUUGAAGAUCG--ACUGGCUG--CCGUUGUUGGCCUUAAAGUCAAAUCCAUG (.(..((((..((.((...((((.-.....))))-...)).))..))))..).).............(--(((....(--(((....)))).....))))......... ( -18.10, z-score = 0.84, R) >consensus CAGCUCCACUGCCUGUUCAACAUCAUGGACCUGCGCCAAUUCGGGGCCACCACCAUAACUGGGGGUCACCACCGGCCGGGUGGUGGUGGGUCGGUGGGUCAGGUCCAUG .......................((((((((((.(((....(.((.((((((((((..((((((....)).))))....)))))))))).)).)..))))))))))))) (-32.00 = -34.08 + 2.08)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:53:25 2011