
| Sequence ID | dm3.chrX |
|---|---|
| Location | 17,411,339 – 17,411,449 |
| Length | 110 |
| Max. P | 0.694698 |

| Location | 17,411,339 – 17,411,449 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 69.22 |
| Shannon entropy | 0.56726 |
| G+C content | 0.36059 |
| Mean single sequence MFE | -22.07 |
| Consensus MFE | -8.33 |
| Energy contribution | -10.30 |
| Covariance contribution | 1.97 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.694698 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
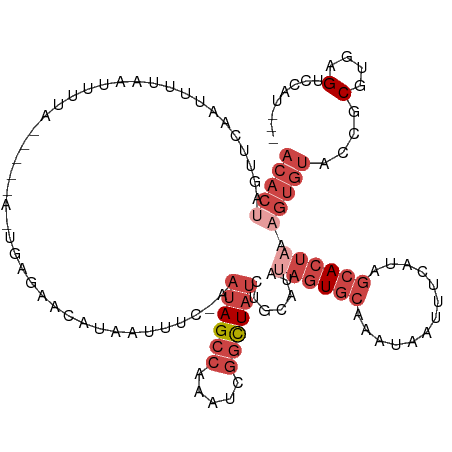
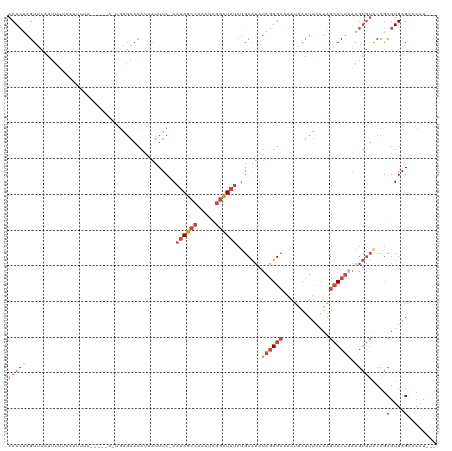
>dm3.chrX 17411339 110 + 22422827 ACACUAGUUCAAUUUUAAUUUUA-----A-UGAGAACAUAAUUUC-AAUAGCCAAAUCGGUUAUCUGCAUAUAGUGCUAAUAAUUUCAUAGCACUAAAGUGUACCGCGUGAGUCCAU--- (((((.((((.(((........)-----)-)..))))........-.((((((.....)))))).......((((((((.........)))))))).)))))...((....))....--- ( -22.70, z-score = -2.44, R) >droAna3.scaffold_13248 1815921 87 - 4840945 -CAAAAGUCAGAAA------------------AGCCCAUUGCUUCGAAAAGGAAAACAAACUGAAGGCGAAAAAUG-----GAUUUUAUGGAACGAGAA---GCCACGUGAGAG------ -......(((.(((------------------(..(((((..((((...((.........)).....)))).))))-----).)))).)))........---....(....)..------ ( -12.20, z-score = 0.32, R) >droEre2.scaffold_4690 7750380 99 + 18748788 ACACUAGUGCAAUU--------------------AAAAUCAUUUC-AAUAGCCAAAUCGGCUAUCUGCAUAUAGUGCAGAUCAUUUUAUAGCACUAAAGUGUGCCGCGUGAGGGCAUUAU ((((((((((...(--------------------(((((......-..(((((.....)))))(((((((...)))))))..))))))..)))))..)))))(((.(....))))..... ( -34.50, z-score = -4.91, R) >droYak2.chrX 16039726 119 + 21770863 ACACCAGUUCAAUUUAAAAGCACCGCACAGUGAGAACAUAAUUUC-AAUAGCCAAAUUGGCUAUCUGCAUAUAGUGCAAACAAUUUUAUGGCACUAAAGUGUACCGCGUGAGAGCAUUAU ......((((...(((...((...((((((((....(((((....-.((((((.....)))))).(((((...))))).......))))).))))...))))...)).)))))))..... ( -25.50, z-score = -1.13, R) >droSec1.super_17 1256921 115 + 1527944 ACACUAGUUCAAUUUUAAUUUUAAUCACA-UGAGAACAUUAUUUC-AAUAGCCAAAUCGGUUAUCUGCAUAUAGUGCCAAUAAUUUCAUAGCACUAAAGUGUACCGCGUGAGUCCAU--- ........................((((.-(((((......))))-)((((((.....)))))).(((((.((((((.............))))))..)))))....))))......--- ( -19.32, z-score = -1.30, R) >droSim1.chrX 3537480 115 - 17042790 ACACUAGUUCAAUUUUAAUUUUAAUCACA-UGAGAACAUAAUUUC-AAUAGCCAAAUCGGUUAUCUGCAUAUAGUGCCAAUAAUUUCAUAGCACUCAAGUGUACCGCGUGAGUCCAU--- ........................((((.-(((((......))))-)((((((.....)))))).(((((..(((((.............)))))...)))))....))))......--- ( -18.22, z-score = -0.97, R) >consensus ACACUAGUUCAAUUUUAAUUUUA_____A_UGAGAACAUAAUUUC_AAUAGCCAAAUCGGCUAUCUGCAUAUAGUGCAAAUAAUUUCAUAGCACUAAAGUGUACCGCGUGAGUCCAU___ .(((...........................................((((((.....)))))).(((((..(((((.............)))))...)))))....))).......... ( -8.33 = -10.30 + 1.97)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:53:04 2011