| Sequence ID | dm3.chrX |
|---|---|
| Location | 17,351,427 – 17,351,579 |
| Length | 152 |
| Max. P | 0.936106 |

| Location | 17,351,427 – 17,351,579 |
|---|---|
| Length | 152 |
| Sequences | 5 |
| Columns | 166 |
| Reading direction | reverse |
| Mean pairwise identity | 75.82 |
| Shannon entropy | 0.41935 |
| G+C content | 0.31671 |
| Mean single sequence MFE | -28.22 |
| Consensus MFE | -19.48 |
| Energy contribution | -19.96 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.40 |
| SVM RNA-class probability | 0.936106 |
| Prediction | RNA |
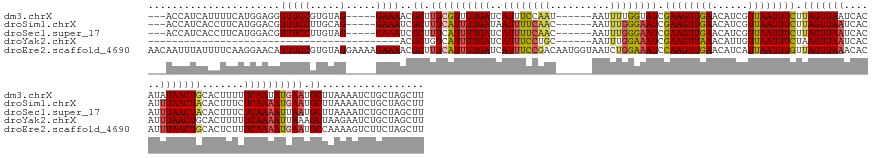
Download alignment: ClustalW | MAF
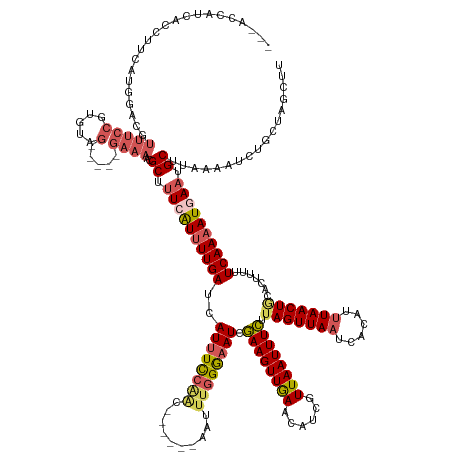
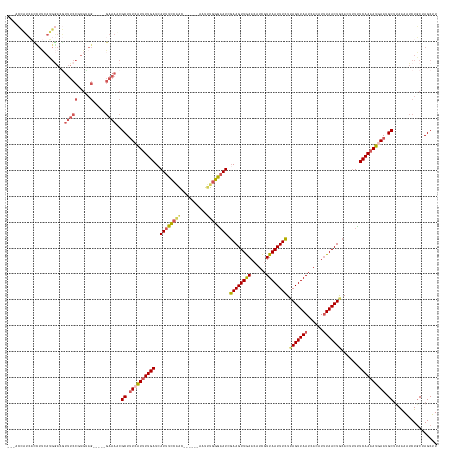
>dm3.chrX 17351427 152 - 22422827 ---ACCAUCAUUUUCAUGGAGGUUUCCGUGUAG-----GAAAACGCUUUCGUUUUGAUCAUUUCCAAU------AAUUUUGGUAUCGAAGUUGAACAUCGUUAAUUUCUUAGUUAAUCACAUAUAACUGCACUUUUUCAAUAUGAAUGCUUAAAAUCUGCUAGCUU ---.((..(((....)))..))(((((.....)-----))))..((.(((((.((((......((((.------....))))....((((((((......)))))))).((((((........)))))).......)))).))))).))................. ( -26.10, z-score = -0.19, R) >droSim1.chrX 3478779 152 + 17042790 ---ACCAUCACCUUCAUGGACGUUUCCUUGCAG-----GAAAUCGCUUUCAUUUUGAUAAUUUUCAAC------AAUUUGGGAAUCGAAGUUGAACAUCGUUAAUUUCUUAGUUAAUCACAUUUAACUACACUUUCUCAAAAUGAAUGCUUAAAAUCUGCUAGCUU ---.((((.......)))).......((.((((-----(.....((.((((((((((..(((..(((.------...)))..))).((((((((......)))))))).(((((((......))))))).......)))))))))).))......))))).))... ( -33.00, z-score = -3.27, R) >droSec1.super_17 1197061 152 - 1527944 ---ACCAUCACCUUCAUGGACGUUUCCUUGUAG-----GAAAUCGCUUUCAUUUUGAUCAUUUUCAAC------AAUUUGGGAAUCGAAGUUGAACAUCGUUAAUUUCUUAGUUAAUCACAUUUAACUACACUUUCUCAAAAUUAAUGCUUAAAAUCUGCUAGCUU ---.......(((.((.(((....))).)).))-----).....(((..((((((((.((((......------..((((((((..((((((((......)))))))).(((((((......)))))))....))))))))...)))).))))))..))..))).. ( -29.60, z-score = -2.69, R) >droYak2.chrX 15979422 118 - 21770863 ------------------------------------------ACGCUGUCAUUUUGAUCAUUUCCUGC------AAUUUGGAAAUCGAAGUUAAACAUUGUUAAUUUCUAAGUUAAUCACAUUUAACUGCACUUUUUCAAAAUUAAAGCUAAGAAUCUGCUAGCUU ------------------------------------------..(((...(((((((..((((((...------.....)))))).((((((((......))))))))..((((((......))))))........)))))))...)))................. ( -21.60, z-score = -1.41, R) >droEre2.scaffold_4690 7690945 166 - 18748788 AACAAUUUAUUUUCAAGGAACAUUUCCGUGUAGGAAAAGAAAACGCUUUCAUUUUGAUCAUUUCCGACAAUGGUAAUCUGGAAAUCCAAGUUGAACAUCAUUAAUUUGUUAGUUAAACACAUUUAACUGCACUCUUUCAAAAUGAAUGCCAAAAGUCUUCUAGCUU ...............((((((.(((((.....))))).......((.((((((((((..(((((((............))))))).((((((((......)))))))).((((((((....)))))))).......)))))))))).)).....)).))))..... ( -30.80, z-score = -0.93, R) >consensus ___ACCAUCACCUUCAUGGACGUUUCCGUGUAG_____GAAAACGCUUUCAUUUUGAUCAUUUCCAAC______AAUUUGGGAAUCGAAGUUGAACAUCGUUAAUUUCUUAGUUAAUCACAUUUAACUGCACUUUUUCAAAAUGAAUGCUUAAAAUCUGCUAGCUU ............................................((.((((((((((..((((((((..........)))))))).((((((((......)))))))).(((((((......))))))).......)))))))))).))................. (-19.48 = -19.96 + 0.48)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:52:47 2011