| Sequence ID | dm3.chrX |
|---|---|
| Location | 17,295,962 – 17,296,058 |
| Length | 96 |
| Max. P | 0.973390 |

| Location | 17,295,962 – 17,296,058 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 61.61 |
| Shannon entropy | 0.70120 |
| G+C content | 0.69515 |
| Mean single sequence MFE | -45.75 |
| Consensus MFE | -19.35 |
| Energy contribution | -21.05 |
| Covariance contribution | 1.70 |
| Combinations/Pair | 1.65 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.89 |
| SVM RNA-class probability | 0.973390 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chrX 17295962 96 + 22422827 CGCCUGGCCGUAUGCGGCCGCUGCCGGCGCCGCC---------------------CACGGAGCCACGCCCCACACGAACAUAGACAUCUAUUACCGUCAAGCGGCCGCUGCGGCCGC .....(((((((.(((((((((...(((.(((..---------------------..))).))).........(((...((((....))))...)))..)))))))))))))))).. ( -48.30, z-score = -3.38, R) >droSim1.chrX 3423330 96 - 17042790 CGCCUGGCCGUAUGCGGCCGCUGCCGGCGCCGCCC---------------------ACGGAGCCACGCCCCACACGAACAUAGACAUAUAUUACCGCCAAGCGGCCGCUGCGGCCGC .....(((((((.(((((((((((.(((.(((...---------------------.))).))).((.......))...................))..)))))))))))))))).. ( -45.40, z-score = -2.43, R) >droSec1.super_17 1142813 96 + 1527944 CGCCUGGCCGUAUGCGGCCGCUGCCGGCGCCGCC---------------------CACGGAGCCACGCCCCACACGAACAUAGACAUCUAUUACCGCCAAGCGGCCGCUGCGGCCGC .....(((((((.(((((((((((.(((.(((..---------------------..))).)))...............((((....))))....))..)))))))))))))))).. ( -47.10, z-score = -2.97, R) >droVir3.scaffold_12970 5729216 111 - 11907090 CGCCUGGCCGUAUGCGGCAGCUGCCGGCGCCGGCGCCGCAGCAGCCGCG------CAUGGCGCACCUCCACAUUCCAGCAUCGAUAUCUACUAUCGUCAAGCGGCCGCGGCGGCGGC ((((((((((((((((((.(((((.((((....))))))))).)))(((------(...))))..............))))(((((.....)))))....))))))).))))..... ( -55.80, z-score = -0.89, R) >droGri2.scaffold_14853 9138403 117 + 10151454 CGCAUGGCCGUAUGCGGCUGCUGCCGGCGCUGGCGUCGCAGCAGCUGCUGCUGCUCAUGGCGGACCUUCCCACUCGAGCAUCGAUAUCUACUAUCGACAGGCGGCGGCGGCAGCCGC .(((((....)))))(((((((((((.((((...((((((((....)))))(((((..((.((......)).)).)))))..((((.....))))))).)))).))))))))))).. ( -61.80, z-score = -2.21, R) >anoGam1.chrU 33401536 90 + 59568033 CGCCUGGCCUUAUCCUACUCCU-UCAGGGCCUCC-----------------------UGGA---ACGUCACAAAGUGCGGUUGACGCAUACUAUAGACAUGCAGCUGCAGCAGCUGC ..((.((((((...........-..))))))...-----------------------.)).---..(((.....(((((.....)))))......)))..(((((((...))))))) ( -27.32, z-score = -0.15, R) >apiMel3.Group3 10145134 92 + 12341916 CAUCCCGCCGCGGGACGAUACUGGCCCUACCCG-------------------------AGCGCGCACGCGCUGCCCGCGAACGGCCUCUUCUACCAGCAAGCGUCGGCCGCGGUCAC ....((((((((((.(.....(((......)))-------------------------(((((....))))))))))))...((((.((((.....).)))....)))))))).... ( -34.50, z-score = 0.31, R) >consensus CGCCUGGCCGUAUGCGGCCGCUGCCGGCGCCGCC_____________________CACGGAGCCACGCCCCACACGAACAUAGACAUCUACUACCGACAAGCGGCCGCUGCGGCCGC .....((((((.((((((((((...((((.................................................................)))).)))))))))))))))).. (-19.35 = -21.05 + 1.70)
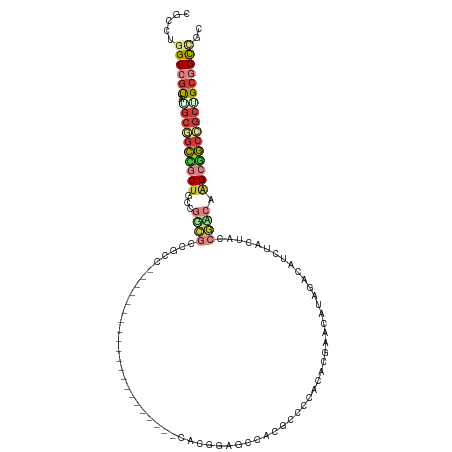


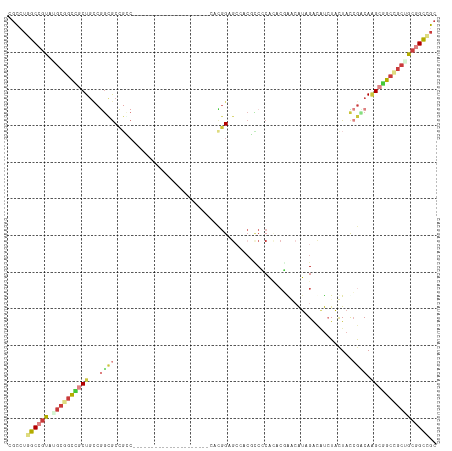
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:52:30 2011