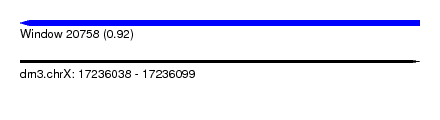
| Sequence ID | dm3.chrX |
|---|---|
| Location | 17,236,038 – 17,236,099 |
| Length | 61 |
| Max. P | 0.917632 |

| Location | 17,236,038 – 17,236,099 |
|---|---|
| Length | 61 |
| Sequences | 5 |
| Columns | 70 |
| Reading direction | reverse |
| Mean pairwise identity | 58.15 |
| Shannon entropy | 0.73948 |
| G+C content | 0.33902 |
| Mean single sequence MFE | -12.32 |
| Consensus MFE | -5.28 |
| Energy contribution | -5.24 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.17 |
| Structure conservation index | 0.43 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.26 |
| SVM RNA-class probability | 0.917632 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
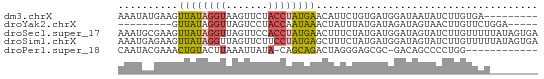
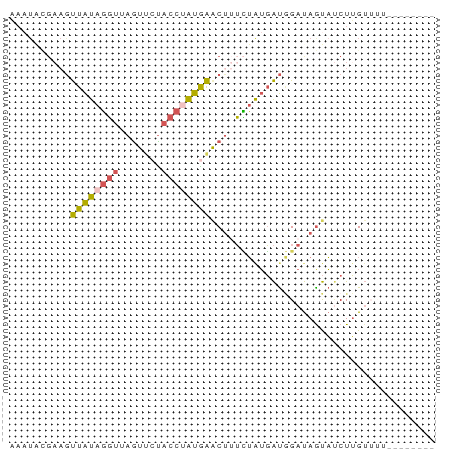
>dm3.chrX 17236038 61 - 22422827 AAAUAUGAAGUUAUAGGUAAGUUCUACCUAUGAACAUUCUGUGAUGGAUAAUAUCUUGUGA--------- ......(((.(((((((((.....)))))))))...)))...((((.....))))......--------- ( -12.10, z-score = -1.45, R) >droYak2.chrX 15852444 56 - 21770863 ---------GUUAUAGGUUAGUCCUACCAAUAAACUAUUUAUGAUAGAUAGUAACUUGUUCUGGA----- ---------....((((.....))))(((((((((((((((...))))))))...))))..))).----- ( -8.90, z-score = -0.10, R) >droSec1.super_17 1082193 70 - 1527944 AAAUGCGAAGUUAUAGGUUAGUUCCACCUAUGAACUUUCUAUGAUGGAUAGUAUCUUGUUUUUAUAGUGA ......((((((((((((.......))))))).)))))((((((.((((((....))))))))))))... ( -12.90, z-score = -0.86, R) >droSim1.chrX 3366733 70 + 17042790 AAAUGAGAAGUUAUAGGUUAGUUCUUCCUAUGAGCUUUCUAUGAUGGAUAGUAUCUUGUUUUUAUAGUGA (((..(((.((((((((..(((((.......))))).))))))))........)))..)))......... ( -13.40, z-score = -0.78, R) >droPer1.super_18 385603 56 - 1952607 CAAUACGAAACUGUACUUAAAUUAUA-CAGCAGACUAGGGAGCGC-GACAGCCCCUGG------------ ..........(((((.........))-)))....((((((.((..-....))))))))------------ ( -14.30, z-score = -2.66, R) >consensus AAAUACGAAGUUAUAGGUUAGUUCUACCUAUGAACUUUCUAUGAUGGAUAGUAUCUUGUUUU________ ..........((((((((.......))))))))..................................... ( -5.28 = -5.24 + -0.04)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:52:18 2011