
| Sequence ID | dm3.chrX |
|---|---|
| Location | 17,117,951 – 17,118,045 |
| Length | 94 |
| Max. P | 0.672354 |

| Location | 17,117,951 – 17,118,045 |
|---|---|
| Length | 94 |
| Sequences | 7 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 64.11 |
| Shannon entropy | 0.70038 |
| G+C content | 0.61876 |
| Mean single sequence MFE | -35.72 |
| Consensus MFE | -19.92 |
| Energy contribution | -21.67 |
| Covariance contribution | 1.75 |
| Combinations/Pair | 1.62 |
| Mean z-score | -0.28 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.672354 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
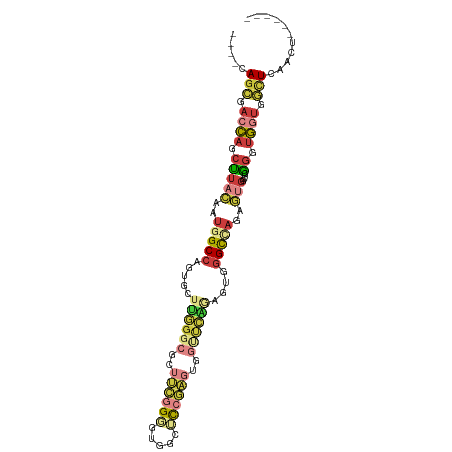
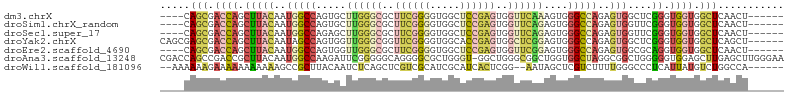
>dm3.chrX 17117951 94 + 22422827 ----CAGCGACCAGCUUACAAUGGCCAGUGCUUGGGCGCUUCGGGGUGGCUCCGAGUGGUUCAAAGUGGGCCAGAGUGGCUCGGGUGGUGGCUCAACU------ ----.(((.((((.(((((..(((((...((((((((..(((((((...)))))))..)))).)))).)))))..)))....)).)))).))).....------ ( -37.50, z-score = -0.29, R) >droSim1.chrX_random 4425815 94 + 5698898 ----CAGCGACCAGCUUACAAUGGCCAGUGCUUGGGCGCUUCGGGGUGGCUCCGAGUGGUUCAGAGUGGGCCAGAGUGGUUCGGGUGGUGGCUCAACU------ ----.(((.((((.(((((..(((((...((((((((..(((((((...)))))))..))))).))).)))))..)))....)).)))).))).....------ ( -37.50, z-score = -0.46, R) >droSec1.super_17 954463 94 + 1527944 ----CAGCGACCAGCUUACAAUGGCCAGAGCUUGGGCGCUUCGGGGUGGCUCCGAGUGGUUCAGAGUGGGCCAGAGUGGUUCGGGUGGUGGCUCAACU------ ----.(((.((((.(((((..(((((...((((((((..(((((((...)))))))..))))).))).)))))..)))....)).)))).))).....------ ( -38.60, z-score = -0.88, R) >droYak2.chrX 15724637 98 + 21770863 CAGCGAGCGACCAGCUUACAAUAGCCAGUGGUUGGGCGGUUCGGGGUGGCACCGAGUGGCUCGGAGUGGGCCAGAGUGGCUCGGGUGGUGGCUCAGCU------ .(((((((.(((((((..((((.(((........))).))).)....))).((((((.((((((......)).)))).)))))).)))).)))).)))------ ( -43.50, z-score = -1.46, R) >droEre2.scaffold_4690 7459633 94 + 18748788 ----CAGCGACCAGCUUACAAUGGCCAGUGGUUGGGCGCUUCGGGGUGGCUCCGAGUGGUUCGGAGUGGGCCAGAGUGGCGCAGGUGGUGGCUCAACU------ ----.(((.((((.((..((((.(....).)))).(((((.(.((.(.((((((((...)))))))).).))...).))))))).)))).))).....------ ( -40.50, z-score = -1.13, R) >droAna3.scaffold_13248 1492392 103 - 4840945 CGACCAGCCGACCGCUUACAAUGGCCAAGAUUCGGGGGCAGGGGCGCUGGGU-GGCUGGGCGGCUGGUGGCUAGGCGGCUGGGGGUGGAGCUUGAGCUUGGGAA ...(((((((.(((((..((.(.(((.((..((.((.((....)).)).)).-..)).))).).))..)))..)))))))))...(.((((....)))).)... ( -40.10, z-score = 0.53, R) >droWil1.scaffold_181096 7573140 94 + 12416693 --AAAAAAGAAAAAAAAAAAGCCGCUUACAAUCUCAGCUCGUCGCAUCGCAUCACUCGG--AAUAGCUCGUCUUUUGGGCCCUCAUUAUGUCUGGCCA------ --..((((((...........(((((.........))).....((...)).......))--.........)))))).((((..(.....)...)))).------ ( -12.35, z-score = 1.74, R) >consensus ____CAGCGACCAGCUUACAAUGGCCAGUGCUUGGGCGCUUCGGGGUGGCUCCGAGUGGUUCAGAGUGGGCCAGAGUGGCUCGGGUGGUGGCUCAACU______ .....(((.((((.(((((..(((((.....((((((..((((((.....))))))..))))))....)))))..)))....)).)))).)))........... (-19.92 = -21.67 + 1.75)


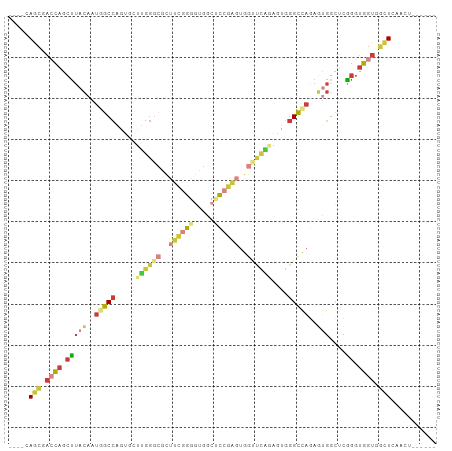
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:51:55 2011