| Sequence ID | dm3.chr2L |
|---|---|
| Location | 11,642,444 – 11,642,513 |
| Length | 69 |
| Max. P | 0.976516 |

| Location | 11,642,444 – 11,642,513 |
|---|---|
| Length | 69 |
| Sequences | 12 |
| Columns | 69 |
| Reading direction | forward |
| Mean pairwise identity | 82.36 |
| Shannon entropy | 0.37925 |
| G+C content | 0.31375 |
| Mean single sequence MFE | -12.84 |
| Consensus MFE | -8.94 |
| Energy contribution | -8.72 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.95 |
| SVM RNA-class probability | 0.976516 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
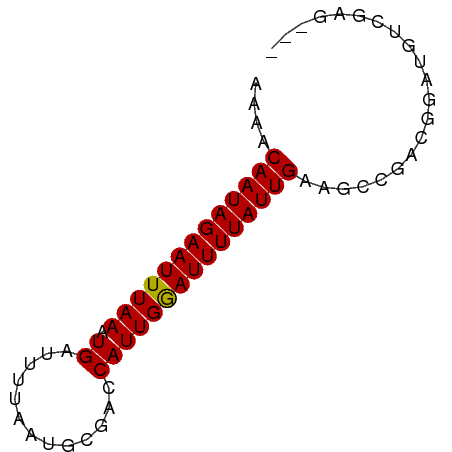
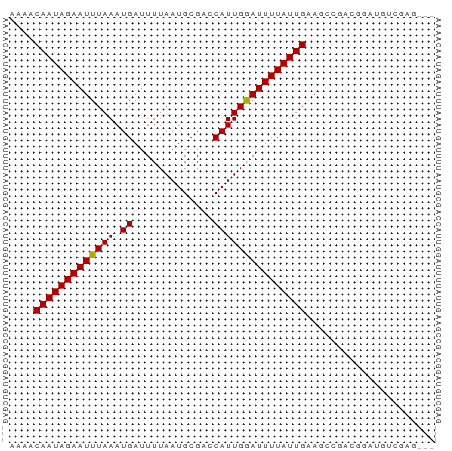
>dm3.chr2L 11642444 69 + 23011544 AAAACAAUAGAAUUUAAAUGAUUUUAAUGCGACCAUUGGAUUUUAUUGAAGCCGACGGAUGUCGAGGCA ....(((((((((((((.((.............)))))))))))))))..((((((....)))..))). ( -14.62, z-score = -1.98, R) >droSim1.chr2L 11450929 69 + 22036055 AAAACAAUAGAAUUUAAAUGAUUUUAAUGCGACCAUUGGAUUUUAUUGAAGCCGACGGAUGUCGAGGCA ....(((((((((((((.((.............)))))))))))))))..((((((....)))..))). ( -14.62, z-score = -1.98, R) >droSec1.super_3 7026620 69 + 7220098 AAAACAAUAGAAUUUAAAUGAUUUUAAUGCGACCAUUGGAUUUUAUUGAAGCCGACGGAUGUCGAGGCA ....(((((((((((((.((.............)))))))))))))))..((((((....)))..))). ( -14.62, z-score = -1.98, R) >droYak2.chr2L 8065368 69 + 22324452 AAAACAAUAGAAUUUAAAUGAUUUUAAUGCGACCAUUGGAUUUUAUUGAAGCCGACGGAUGUCGAGGCA ....(((((((((((((.((.............)))))))))))))))..((((((....)))..))). ( -14.62, z-score = -1.98, R) >droEre2.scaffold_4929 12872431 69 - 26641161 AAAACAAUAGAAUUUAAAUGAUUUUAAUGCGACCAUUGGAUUUUAUUGAAGCCGACGGAUGUCGAGGCA ....(((((((((((((.((.............)))))))))))))))..((((((....)))..))). ( -14.62, z-score = -1.98, R) >droAna3.scaffold_12916 9360672 66 - 16180835 AAAACAAUAGAAUUUAAAUGAUUUUAAUGCGACCAUUGAAUUUUAUUGAAGUCGACGGAUGUCGAG--- ....(((((((((((((.((.............)))))))))))))))...(((((....))))).--- ( -14.52, z-score = -2.53, R) >dp4.chr4_group3 2502703 54 + 11692001 AGAACAAUAGAAUUUAAAUGAUUUUAAUGCGAGCAUUGAAUUUUAUUGAAGGCA--------------- ....(((((((((((((.((.(((......))))))))))))))))))......--------------- ( -9.70, z-score = -2.21, R) >droPer1.super_1 3983197 54 + 10282868 AGAACAAUAGAAUUUAAAUGAUUUUAAUGCGAGCAUUGAAUUUUAUUGAAGGCA--------------- ....(((((((((((((.((.(((......))))))))))))))))))......--------------- ( -9.70, z-score = -2.21, R) >droWil1.scaffold_180772 5087161 69 - 8906247 AAAACAAUAGAAUUUAAAUGAUUUUAAUGCGACCAUUGAAUUUUAUUGAAGCUGCCUUCACCCUGGACG ....(((((((((((((.((.............)))))))))))))))......((........))... ( -10.12, z-score = -1.89, R) >droVir3.scaffold_12963 6202892 62 + 20206255 AAAACAAUAGAAUUUAAAUGAUUUUAAUGCGUACAUUGGAUUUUAUUGAAGCCAACGAGGCA------- ....(((((((((((((.((((........)).)))))))))))))))..(((.....))).------- ( -13.00, z-score = -2.89, R) >droMoj3.scaffold_6500 25334811 62 + 32352404 AAAACAAUAGAAUUUAAAUGAUUUUAAUGCGUACAUUGGAUUUUAUUGAAGCCAACGAGGCA------- ....(((((((((((((.((((........)).)))))))))))))))..(((.....))).------- ( -13.00, z-score = -2.89, R) >droGri2.scaffold_15252 9467953 62 + 17193109 AAAACAAUAGAAUUUAAAUGAUUUUAAUGCGUACAUUGGAUUUUAUUGAAUCCAGCGAGGGU------- ....((.(((((((.....))))))).))(((....(((((((....)))))))))).....------- ( -10.90, z-score = -1.70, R) >consensus AAAACAAUAGAAUUUAAAUGAUUUUAAUGCGACCAUUGGAUUUUAUUGAAGCCGACGGAUGUCGAG___ ....(((((((((((((.((.............)))))))))))))))..................... ( -8.94 = -8.72 + -0.22)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:33:12 2011