
| Sequence ID | dm3.chrX |
|---|---|
| Location | 17,075,334 – 17,075,399 |
| Length | 65 |
| Max. P | 0.793649 |

| Location | 17,075,334 – 17,075,399 |
|---|---|
| Length | 65 |
| Sequences | 5 |
| Columns | 66 |
| Reading direction | reverse |
| Mean pairwise identity | 91.47 |
| Shannon entropy | 0.13919 |
| G+C content | 0.39882 |
| Mean single sequence MFE | -9.52 |
| Consensus MFE | -8.30 |
| Energy contribution | -8.30 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.87 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.793649 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
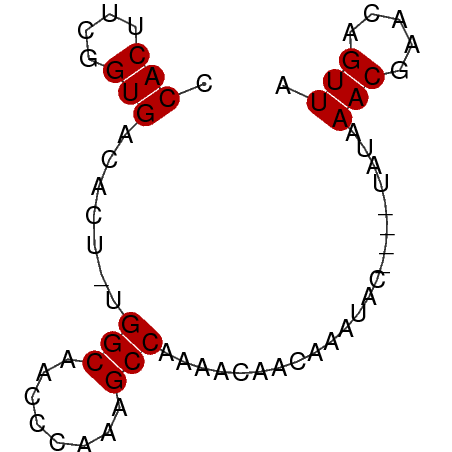
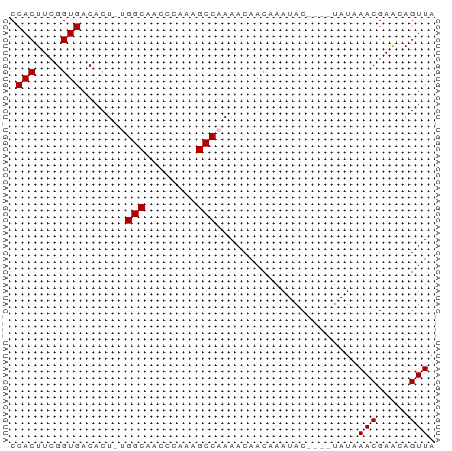
>dm3.chrX 17075334 65 - 22422827 CCACUUCGGUGACACU-UGGCAACCCAAAGCCAAAACAACAAAUACAAAAUAUAAACGAACAGUUA .(((....)))....(-((((........)))))....................(((.....))). ( -11.10, z-score = -2.64, R) >droSim1.chrX 13255623 62 - 17042790 CCACUUCGGUGACACUACGGCAAUCCAAAGCCAAAACAACAAAUAC----UAUAAACGAACAGUUA .(((....))).......(((........)))..............----....(((.....))). ( -9.70, z-score = -2.28, R) >droSec1.super_17 916802 62 - 1527944 CCACUUCGGUGACACUACGGCAAUCCAAAGCCAAAACAACAAAUAC----UAUAAACGAGCAGUUA .(((....))).......(((........)))..............----....(((.....))). ( -9.70, z-score = -1.77, R) >droYak2.chrX 15686467 61 - 21770863 CCACUUUGGUGACACU-UGGCAACCGAAAGCCAAAACAACAAAUAC----UAUUAACGAACAGUUA ......((....)).(-((((........)))))............----...((((.....)))) ( -8.80, z-score = -1.10, R) >droEre2.scaffold_4690 7422099 61 - 18748788 CCACUUUGGUGACACU-UGGCAACCGAAAGCCAAAACAACAAAUAC----UAUAAACGAACAGUUA ......((....)).(-((((........)))))............----....(((.....))). ( -8.30, z-score = -0.95, R) >consensus CCACUUCGGUGACACU_UGGCAACCCAAAGCCAAAACAACAAAUAC____UAUAAACGAACAGUUA .(((....))).......(((........)))......................(((.....))). ( -8.30 = -8.30 + -0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:51:51 2011