
| Sequence ID | dm3.chrX |
|---|---|
| Location | 16,938,339 – 16,938,415 |
| Length | 76 |
| Max. P | 0.627171 |

| Location | 16,938,339 – 16,938,415 |
|---|---|
| Length | 76 |
| Sequences | 11 |
| Columns | 90 |
| Reading direction | forward |
| Mean pairwise identity | 82.82 |
| Shannon entropy | 0.33414 |
| G+C content | 0.35784 |
| Mean single sequence MFE | -17.56 |
| Consensus MFE | -10.30 |
| Energy contribution | -10.30 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.627171 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 16938339 76 + 22422827 --------AGACCCAAAG--UU--CGUUAUUGAUGGCGAAUAUUACGCCAACUAAGCAAAUUAAAUUGUUUACAUUG-ACAAAGG-CCAA --------..........--..--.((((.((.(((((.......)))))..(((((((......))))))))).))-)).....-.... ( -14.70, z-score = -1.66, R) >droSim1.chrX 13116444 76 + 17042790 --------AGACCCAAAG--UU--CUUUAUUGAUGGCGAAUAUUACGCCAACUAAGCAAAUUAAAUUGUUUACAUUG-ACAAAGG-CCAA --------.........(--..--((((.(..((((((.......))))...(((((((......))))))).))..-).)))).-.).. ( -14.80, z-score = -1.98, R) >droSec1.super_17 771609 76 + 1527944 --------AGACCCAAAG--UU--CUUUAUUGAUGGCGAAUAUUACGCCAACUAAGCAAAUUAAAUUGUUUACAUUG-ACAAAGG-CCAA --------.........(--..--((((.(..((((((.......))))...(((((((......))))))).))..-).)))).-.).. ( -14.80, z-score = -1.98, R) >droYak2.chrX 15550158 76 + 21770863 --------AGACCCAAAG--UU--UUUUAUUGAUGGCGAAUAUUACGCCAACUAAGCAAAUUAAAUUGUUUACAUUG-ACAAAGG-CCAA --------.........(--((--(((((.((.(((((.......)))))..(((((((......))))))))).))-)..))))-)... ( -12.20, z-score = -1.01, R) >droEre2.scaffold_4690 7292705 76 + 18748788 --------AGACCCAAAG--UU--CUUUAUUGAUGGCGAAUAUUACGCCAACUAAGCAAAUUAAAUUGUUUACAUUG-ACAAAGG-CCAA --------.........(--..--((((.(..((((((.......))))...(((((((......))))))).))..-).)))).-.).. ( -14.80, z-score = -1.98, R) >droAna3.scaffold_13334 750307 76 - 1562580 --------AGUCCCAAAG--UU--CUUUAUUGAUGGCGAAUAUUACGCCAACUAAGCAAAUUAAAUUGUUUACAUUG-ACAAAGG-CCAA --------.........(--..--((((.(..((((((.......))))...(((((((......))))))).))..-).)))).-.).. ( -14.80, z-score = -1.69, R) >dp4.chrXL_group1e 1072306 80 + 12523060 --------ACACUCGUGG--UCUCCUUUAUUGAUGGCGAAUAUUACGCCAACUAAGCAAAUUAAAUUGUUUACAUUGGACAAAGGGCCAA --------.......(((--(((..((((.((.(((((.......)))))..(((((((......))))))))).))))....)))))). ( -18.60, z-score = -1.73, R) >droPer1.super_30 834006 80 - 958394 --------ACACUCGUGG--UCUCCUUUAUUGAUGGCGAAUAUUACGCCAACUAAGCAAAUUAAAUUGUUUACAUUGGACAAAGGGCCAA --------.......(((--(((..((((.((.(((((.......)))))..(((((((......))))))))).))))....)))))). ( -18.60, z-score = -1.73, R) >droVir3.scaffold_12970 9648366 88 + 11907090 GCCCCAAAAGUUUUGUGCGCUACUUUUUAUUGAUGGCGAAUAUUACGCCAACUAAGCAAAUUAAAUUGUUUACAUUG-GCUAAGGGCAA- ((((.....((..(((.((((((........).))))).)))..))(((((.(((((((......)))))))..)))-))...))))..- ( -27.10, z-score = -3.14, R) >droMoj3.scaffold_6473 5019452 84 - 16943266 UCCCCAAAAAACUUGCG---UACUUUUUAUUGAUGGCGAAUAUUACGCCAACUAAGCAAAUUAAAUUGUUUACAUUG-GCAAAGGGCA-- ..(((.......((((.---...............)))).......(((((.(((((((......)))))))..)))-))...)))..-- ( -16.89, z-score = -0.88, R) >droGri2.scaffold_14853 9768894 85 - 10151454 AGCCCCAAAGUUUUGUGUGCUACAUUUUAUUGAUGGCGAAUAUUACGCCAACUAAGCAAAUUAAAUUGUUUACAUUG-GC-AAGGGC--- .((((....((..(((.(((((((......)).))))).)))..))(((((.(((((((......)))))))..)))-))-..))))--- ( -25.90, z-score = -2.86, R) >consensus ________AGACCCAAAG__UU__CUUUAUUGAUGGCGAAUAUUACGCCAACUAAGCAAAUUAAAUUGUUUACAUUG_ACAAAGG_CCAA .................................(((((.......)))))..(((((((......))))))).................. (-10.30 = -10.30 + -0.00)

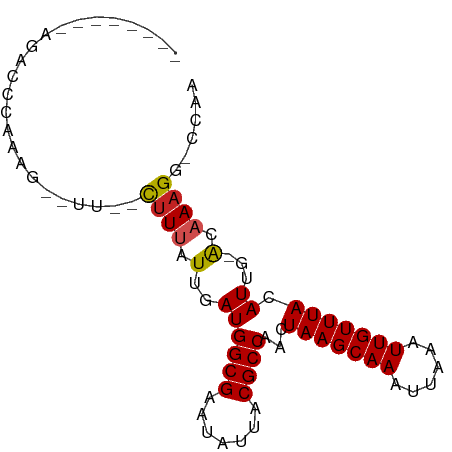
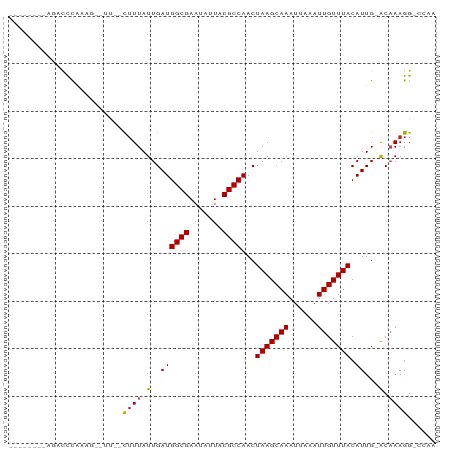
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:51:13 2011