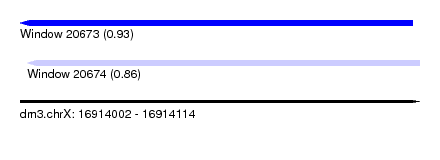

| Sequence ID | dm3.chrX |
|---|---|
| Location | 16,914,002 – 16,914,114 |
| Length | 112 |
| Max. P | 0.927926 |

| Location | 16,914,002 – 16,914,112 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 91.47 |
| Shannon entropy | 0.14848 |
| G+C content | 0.47705 |
| Mean single sequence MFE | -36.83 |
| Consensus MFE | -29.38 |
| Energy contribution | -28.74 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.45 |
| Structure conservation index | 0.80 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.33 |
| SVM RNA-class probability | 0.927926 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
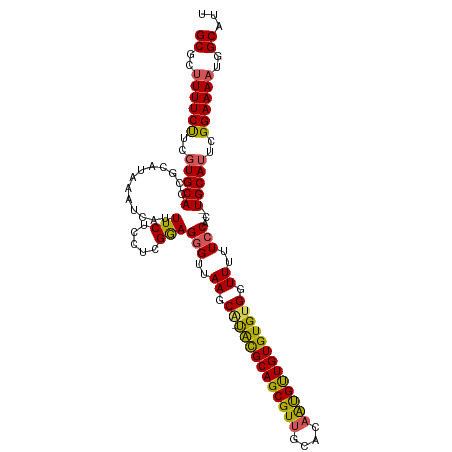
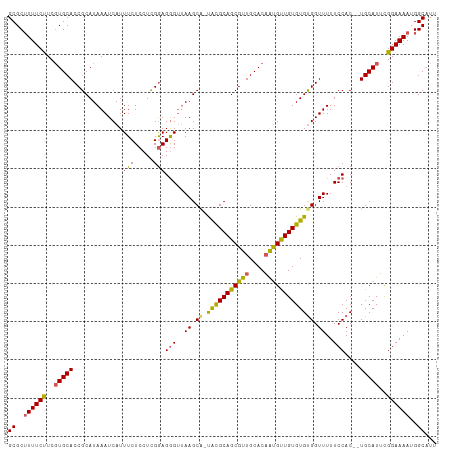
>dm3.chrX 16914002 110 - 22422827 GCGCUUUUCCUCGUGCACCGCAUAAAUCAUUUCUCCUCGAAGGGAGAAGCGUUGUGCAGCGUUGCACAAUGUUGUGUGUGGUUUUUCCAC--UGCAUUCGGAAAAUGGCAUU ((..((((((..(((((..(((((......(((((((....)))))))(((((((((......))))))))))))))((((.....))))--)))))..))))))..))... ( -42.50, z-score = -3.71, R) >droEre2.scaffold_4690 7268222 108 - 18748788 GCGC-UUUCUUCUUGCACCGCAUAAAUCAUUUCUCCUUGGAGGGUUAAGCA-UACGCAGCGUUGCACAGUGCUGUGUGUGGUUUUUCCAC--UGCAUUUGGAAAAUGGCAUU ((((-.........))...........(((((.(((.((.((((..((.((-(((((((((((....))))))))))))).))...)).)--).))...))))))))))... ( -33.20, z-score = -1.32, R) >droYak2.chrX 15523711 111 - 21770863 GCGCUUUUCCUCGUGCACCGCAUAAAUCAUUUCUCCUUGGAGAGUUAAGCA-UACGCAGCGUUGCACCGCGUUGUGUGCGGUUUUUCCACAUUGCAUUCGGAAAACGGCAUU ((..((((((..(((((((((.........(((((....))))).......-.(((((((((......))))))))))))))...........))))..))))))..))... ( -35.50, z-score = -1.87, R) >droSec1.super_17 747539 109 - 1527944 GCGCUUUUCUUCGUGCACCGCAUAAAUCAUUUCUCCUCGGAGGGUUAAGCA-CACGCAGCGUUGCACAAUGUUGUGUGUGGUUUUUCCAC--UGCAUUCGGAAAAUGGCAUU ((..((((((..((((((((.................))).(((..((.((-(((((((((((....))))))))))))).))..)))..--)))))..))))))..))... ( -37.43, z-score = -2.89, R) >droSim1.chrX 13095585 109 - 17042790 GCGCUUUUCUUCGUGCACCGCAUAAAUCAUUUCUCCUCGGAGGGUUAAGCA-UACGCAGCGUUGCACAAUGUUGUGUGUGGUUUUUCCAC--UGCAUUCGGAAAAUGGCAUU ((..((((((..((((((((.................))).(((..((.((-(((((((((((....))))))))))))).))..)))..--)))))..))))))..))... ( -35.53, z-score = -2.45, R) >consensus GCGCUUUUCUUCGUGCACCGCAUAAAUCAUUUCUCCUCGGAGGGUUAAGCA_UACGCAGCGUUGCACAAUGUUGUGUGUGGUUUUUCCAC__UGCAUUCGGAAAAUGGCAUU ((..((((((..(((((((((.........(((((....))))).........((((((((((....)))))))))))))))...........))))..))))))..))... (-29.38 = -28.74 + -0.64)

| Location | 16,914,004 – 16,914,114 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 91.47 |
| Shannon entropy | 0.14848 |
| G+C content | 0.49533 |
| Mean single sequence MFE | -37.13 |
| Consensus MFE | -29.68 |
| Energy contribution | -29.04 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.80 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.864671 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
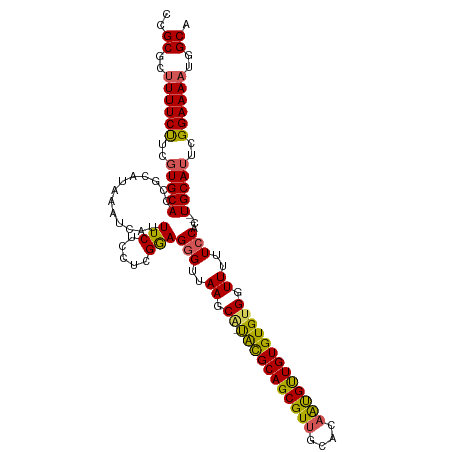
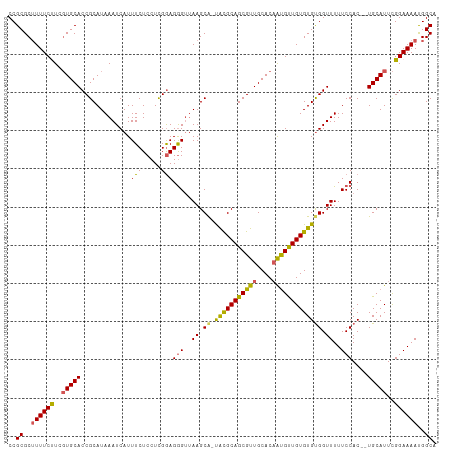
>dm3.chrX 16914004 110 - 22422827 CCGCGCUUUUCCUCGUGCACCGCAUAAAUCAUUUCUCCUCGAAGGGAGAAGCGUUGUGCAGCGUUGCACAAUGUUGUGUGUGGUUUUUCCAC--UGCAUUCGGAAAAUGGCA ..((..((((((..(((((..(((((......(((((((....)))))))(((((((((......))))))))))))))((((.....))))--)))))..))))))..)). ( -42.80, z-score = -3.53, R) >droEre2.scaffold_4690 7268224 108 - 18748788 CCGCGC-UUUCUUCUUGCACCGCAUAAAUCAUUUCUCCUUGGAGGGUUAAGCA-UACGCAGCGUUGCACAGUGCUGUGUGUGGUUUUUCCAC--UGCAUUUGGAAAAUGGCA ..((((-.........))...........(((((.(((.((.((((..((.((-(((((((((((....))))))))))))).))...)).)--).))...)))))))))). ( -33.50, z-score = -1.11, R) >droYak2.chrX 15523713 111 - 21770863 CCGCGCUUUUCCUCGUGCACCGCAUAAAUCAUUUCUCCUUGGAGAGUUAAGCA-UACGCAGCGUUGCACCGCGUUGUGUGCGGUUUUUCCACAUUGCAUUCGGAAAACGGCA ..((..((((((..(((((((((.........(((((....))))).......-.(((((((((......))))))))))))))...........))))..))))))..)). ( -35.80, z-score = -1.69, R) >droSec1.super_17 747541 109 - 1527944 CCGCGCUUUUCUUCGUGCACCGCAUAAAUCAUUUCUCCUCGGAGGGUUAAGCA-CACGCAGCGUUGCACAAUGUUGUGUGUGGUUUUUCCAC--UGCAUUCGGAAAAUGGCA ..((..((((((..((((((((.................))).(((..((.((-(((((((((((....))))))))))))).))..)))..--)))))..))))))..)). ( -37.73, z-score = -2.61, R) >droSim1.chrX 13095587 109 - 17042790 CCGCGCUUUUCUUCGUGCACCGCAUAAAUCAUUUCUCCUCGGAGGGUUAAGCA-UACGCAGCGUUGCACAAUGUUGUGUGUGGUUUUUCCAC--UGCAUUCGGAAAAUGGCA ..((..((((((..((((((((.................))).(((..((.((-(((((((((((....))))))))))))).))..)))..--)))))..))))))..)). ( -35.83, z-score = -2.19, R) >consensus CCGCGCUUUUCUUCGUGCACCGCAUAAAUCAUUUCUCCUCGGAGGGUUAAGCA_UACGCAGCGUUGCACAAUGUUGUGUGUGGUUUUUCCAC__UGCAUUCGGAAAAUGGCA ..((..((((((..(((((((((.........(((((....))))).........((((((((((....)))))))))))))))...........))))..))))))..)). (-29.68 = -29.04 + -0.64)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:51:10 2011