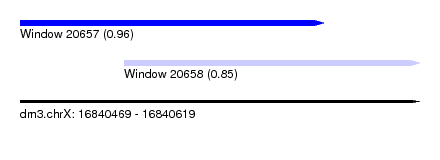

| Sequence ID | dm3.chrX |
|---|---|
| Location | 16,840,469 – 16,840,619 |
| Length | 150 |
| Max. P | 0.955744 |

| Location | 16,840,469 – 16,840,583 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 77.71 |
| Shannon entropy | 0.40076 |
| G+C content | 0.39888 |
| Mean single sequence MFE | -32.50 |
| Consensus MFE | -19.32 |
| Energy contribution | -20.80 |
| Covariance contribution | 1.48 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.62 |
| SVM RNA-class probability | 0.955744 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
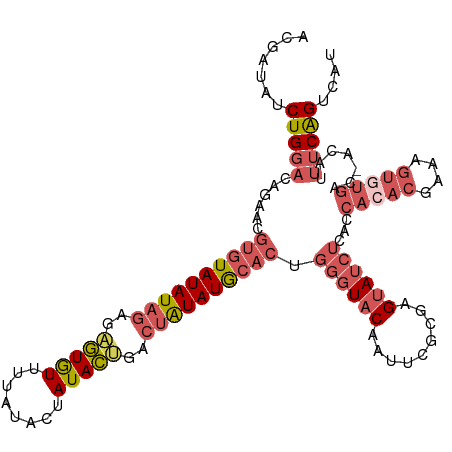
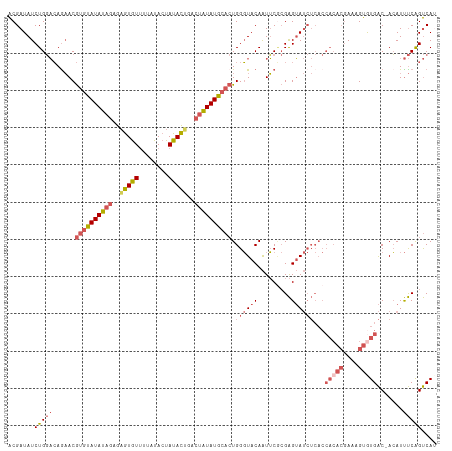
>dm3.chrX 16840469 114 + 22422827 ACGAUAUCUGGACAGAACGUGUAUAUAGAGAGUGUUUUAUACUAUACUAUCUAUAUGCACUGGGUACAAUUCACGAGUAUCUCACCACACGAAAGUGUGAC-ACAUUUCGGUCAA ..........(((.(((.(((((((((((.(((((........))))).))))))))))).((((((.........))))))...(((((....)))))..-....))).))).. ( -36.20, z-score = -3.54, R) >droSim1.chrX_random 4379144 114 + 5698898 ACGAUAUCUGGACAGAACGUGUAUAUAGAGAGUGUUUUAUACUAUACUGACUAUAUGCACUGGGUACAAUUCGCGAGUAUCUCACCACACGAAAGUGUGAC-ACAUUUCAGUCAU ..........(((.(((.((((((((((..(((((........)))))..)))))))))).((((((.........))))))...(((((....)))))..-....))).))).. ( -33.30, z-score = -2.38, R) >droSec1.super_17 674018 114 + 1527944 ACGAUAUCUGGACAGAACGUGUAUAUAGAGAGUGUUUUAUACUAUACUGACUAUAUGCACUGGGUACAAUUCGCGAGUAUCUCACCACACGAAAGUGUGAC-ACAUUUCAGUCAU ..........(((.(((.((((((((((..(((((........)))))..)))))))))).((((((.........))))))...(((((....)))))..-....))).))).. ( -33.30, z-score = -2.38, R) >droYak2.chrX 15450508 114 + 21770863 ACGAUAUCUGGACAAAACGUGUAUAUAGAGAGUGUUUUAUACUAUACGAGCUAUAUGCACUGGGUACAAUUCGCGAGUAUCUCACCACACGAAAGUAUGAC-ACAUUUCGGUCAU ........(((.......((((((((((...((((........))))...)))))))))).((((((.........))))))..))).((....))(((((-........))))) ( -27.70, z-score = -1.21, R) >droAna3.scaffold_13334 981590 99 - 1562580 GCGAUAUCUGCCGAGCACAUAUAUAUUAUAUAUAUCAC-UACUAUAUAGUGUGUAUA---UGGCUACAGUGCCAGGGGA-----------AAGUGUAUGGCCAGAUCCCAGCCA- ((...((((((((.((((((((((((((((((......-...)))))))))))))).---((((......)))).....-----------..)))).))).)))))....))..- ( -32.00, z-score = -1.67, R) >consensus ACGAUAUCUGGACAGAACGUGUAUAUAGAGAGUGUUUUAUACUAUACUGACUAUAUGCACUGGGUACAAUUCGCGAGUAUCUCACCACACGAAAGUGUGAC_ACAUUUCAGUCAU ..........(((.....((((((((((..(((((........)))))..))))))))))..............(((........(((((....))))).......))).))).. (-19.32 = -20.80 + 1.48)

| Location | 16,840,508 – 16,840,619 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 70.77 |
| Shannon entropy | 0.52533 |
| G+C content | 0.40276 |
| Mean single sequence MFE | -26.62 |
| Consensus MFE | -12.24 |
| Energy contribution | -15.00 |
| Covariance contribution | 2.76 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.854248 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
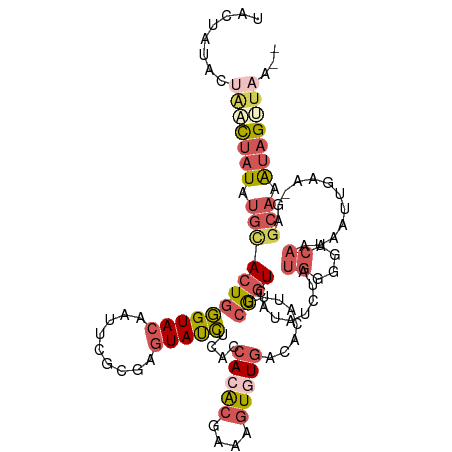
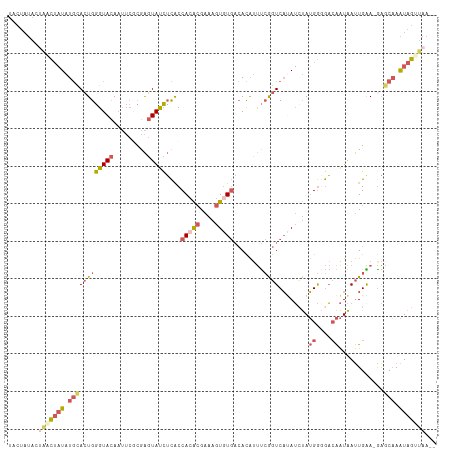
>dm3.chrX 16840508 111 + 22422827 UACUAUACUAUCUAUAUGCACUGGGUACAAUUCACGAGUAUCUCACCACACGAAAGUGUGACACAUUUCGGUCAAAUCUAUGGUGACAAUAAUUGAA-GAGCAAAUAGUUAA-- ...........((((.(((.((.(((((.........)))))(((((((((....)))((((........))))......))))))..........)-).))).))))....-- ( -25.20, z-score = -1.74, R) >droSim1.chrX_random 4379183 111 + 5698898 UACUAUACUGACUAUAUGCACUGGGUACAAUUCGCGAGUAUCUCACCACACGAAAGUGUGACACAUUUCAGUCAUAUCUAUGGGGACAAUAAUUGAA-GAGCAAAUAGUUAA-- ........(((((((.((((((((((((.........)))))....(((((....)))))........))))........((....)).........-..))).))))))).-- ( -28.40, z-score = -2.48, R) >droSec1.super_17 674057 111 + 1527944 UACUAUACUGACUAUAUGCACUGGGUACAAUUCGCGAGUAUCUCACCACACGAAAGUGUGACACAUUUCAGUCAUAUCUAUGGGGACAAUAAUUGAA-GAGCAAAUAGUUAA-- ........(((((((.((((((((((((.........)))))....(((((....)))))........))))........((....)).........-..))).))))))).-- ( -28.40, z-score = -2.48, R) >droYak2.chrX 15450547 113 + 21770863 UACUAUACGAGCUAUAUGCACUGGGUACAAUUCGCGAGUAUCUCACCACACGAAAGUAUGACACAUUUCGGUCAUAUUUAUGGGGACAAUAAUUGGA-AAACAAAUAGCUAAAC .........((((((.((....((((((.........))))))..(((.....(((((((((........))))))))).((....)).....))).-...)).)))))).... ( -24.10, z-score = -2.27, R) >droAna3.scaffold_13334 981628 93 - 1562580 -----------UACUAUAUAGUGUGUAUA-----UGGCUACAGUGCCAGGGGAAAGUGU---AUGGCCAGAUCCCAGCCAGGAAAUAGGUGACUGGAUGGGCCAGUGGGGAG-- -----------..(((((((.(.......-----((((......))))......).)))---)))).....((((.(((........))).(((((.....))))).)))).-- ( -27.02, z-score = 0.09, R) >consensus UACUAUACUAACUAUAUGCACUGGGUACAAUUCGCGAGUAUCUCACCACACGAAAGUGUGACACAUUUCGGUCAUAUCUAUGGGGACAAUAAUUGAA_GAGCAAAUAGUUAA__ ........(((((((.((((((((((((.........)))))....(((((....)))))........))))........((....))............))).)))))))... (-12.24 = -15.00 + 2.76)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:50:57 2011