
| Sequence ID | dm3.chrX |
|---|---|
| Location | 16,329,597 – 16,329,705 |
| Length | 108 |
| Max. P | 0.680545 |

| Location | 16,329,597 – 16,329,705 |
|---|---|
| Length | 108 |
| Sequences | 11 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 76.94 |
| Shannon entropy | 0.48484 |
| G+C content | 0.50229 |
| Mean single sequence MFE | -34.66 |
| Consensus MFE | -16.75 |
| Energy contribution | -17.05 |
| Covariance contribution | 0.30 |
| Combinations/Pair | 1.58 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.680545 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 16329597 108 + 22422827 CGACCUUUCUGCCCCUUCCAG--UGGGGCUGUGAUCUGUGCGGCGAGUAUUUGUCGCGCAAGGAGAAGCUAAUGAAUCACAUUAACAACCAUUUGAAGGAGGAGGUCAUA------ .(((((.....(((((((.((--(((((((......((((((((((....))))))))))......))))((((.....)))).....))))).))))).)))))))...------ ( -40.40, z-score = -2.89, R) >droSec1.super_17 149347 108 + 1527944 CGACCUUUCUGCCCCUUCCAG--UGGGGCUGUGAUCUGUGCGGCGAGUAUCUGUCGCGCAAGGAGAAGCUGAUGAACCAUAUUAACAACCAUUUGAAGGAGGAUGUCAUA------ .(((...(((...(((((.((--(((((((......(((((((((......)))))))))......)))).(((...)))........))))).))))).))).)))...------ ( -33.90, z-score = -0.99, R) >droYak2.chrX 10449003 108 + 21770863 CGACCCUUCUGCCCCUUCCAG--UGGGGCUGCGAUCUGUGCGGCGAGUAUCUGUCGCGCAAGGAGAAGCUGAUGAACCACAUUAAUAACCAUUUAAAGGAGGAGGUCAUA------ .((((((((((((((......--.))))).((..(((((((((((......))))))))..)))...))(((((.....))))).............))))).))))...------ ( -39.30, z-score = -2.17, R) >droEre2.scaffold_4690 7980736 108 - 18748788 CGACCCUUCUGCCCCUUCCAG--UGGGGCUGCGAUCUAUGCGGCGAGUAUCUGUCGCGCAAGGAGAAGCUGAUGAACCACAUUAACAACCAUUUGAAGGAGGAGGUCAUA------ .((((.((((...(((((.((--(((((((....(((.((((.(((.......))))))).)))..))))((((.....)))).....))))).)))))))))))))...------ ( -37.40, z-score = -1.57, R) >droAna3.scaffold_13335 1484023 108 + 3335858 CGACCGUUCUGCCCCUUCCAG--UGGGGCUGUGAUCUGUGCGGCGAGUAUCUAUCCCGCAAGGAGAAGCUCAUGAAUCACAUCAACAAUCAUCUCAAGGAGGAGGUGAUU------ .....(((..(((((......--.)))))((((((..(((.((((.....)..((((....)).)).))))))..))))))..)))(((((((((......)))))))))------ ( -37.40, z-score = -1.35, R) >dp4.chrXL_group1e 9426271 108 - 12523060 CGACCAUUUUGUCCCUUCCAA--UGGGGCUGUGACAUGUGCGGCGAGUACUUGUCGCGCAAGGAGAAGCUCAUGAAUCACAUCAAUAAUCAUUUGAAGGAGGAUGUCAUU------ .(((.(((((...(((((.((--(((((((.(..(.((((((((((....)))))))))).)..).))))).(((......))).....)))).)))))))))))))...------ ( -36.20, z-score = -2.14, R) >droPer1.super_17 1899675 108 - 1930428 CGACCAUUUUGUCCCUUUCAA--UGGGGCUGUGACAUGUGCGGCGAGUACUUGUCGCGCAAGGAGAAGGUCAUGAAUCACCUCAAUAAUCAUUUGAAGGAGGAUGUCAUU------ .((((.(((((((((......--.))))).....(.((((((((((....)))))))))).))))).))))(((((((.((((((.......))).)))..))).)))).------ ( -37.90, z-score = -2.60, R) >droWil1.scaffold_181096 5725975 108 + 12416693 CGUCCUUUUUGUCCCUUUCAA--UGGGGAUGUGAUCUAUGUGGCGAAUACUUGUCACGCAAAGAGAAGCUCAUCAAUCAUAUUAACAAUCACCUUAAGGAGGAUGUUAUA------ ((((((((((((((((.....--.))))))(((((....(((((((....)))))))((........))..................)))))...)))))))))).....------ ( -30.90, z-score = -2.44, R) >droVir3.scaffold_12472 582863 114 - 763072 CGUCCAUUCUGUCCAUUCCAA--UGGGGCUGUGAUCUGUGCGGCGAGUACUUGUCGCGCAAGGAGAAGCUCAUCAAUCACAUUAACAAUCAUCUCAAAGAGGAUGUAAUUGUACCU (((((.....((((.......--..))))((((((.((((((((((....))))))))))..(((...)))....))))))...................)))))........... ( -30.30, z-score = -0.71, R) >droMoj3.scaffold_6359 2692965 101 + 4525533 CGCCCUCCUCAUCCAGUUCAUCCUCCAGCU-CGUUCUGCUCGAGCAGCAACAAUUCGGCU--GCCACGUCCGCCAGCCUGAGCAGC--UCGUCGGCCACGCGCUCU---------- ...........................(((-((((((((....)))).))).....((((--(.(......).))))).))))(((--.(((.....))).)))..---------- ( -26.90, z-score = -0.39, R) >droGri2.scaffold_14853 6839143 108 + 10151454 CGUCCAUUCUGUCCAUUCCAA--UGGGGCUGUGAUCUGUGCGGCGAGUAUUUGUCGCGCAAGGAGAAGCUCAUCAAUCACAUUAACAAUCAUCUCAAGGAGGAUGUGAUU------ (((((.((((((((.......--..))))((((((.((((((((((....))))))))))..(((...)))....))))))...............))))))))).....------ ( -30.70, z-score = -0.61, R) >consensus CGACCAUUCUGUCCCUUCCAA__UGGGGCUGUGAUCUGUGCGGCGAGUAUUUGUCGCGCAAGGAGAAGCUCAUGAAUCACAUUAACAAUCAUUUGAAGGAGGAUGUCAUA______ ....((((((((((...........)))).......(((((((((......))))))))).......................................))))))........... (-16.75 = -17.05 + 0.30)


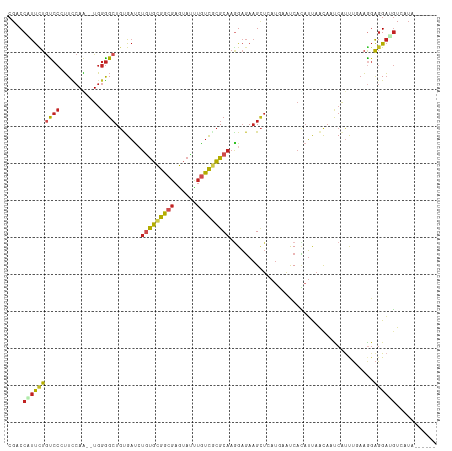
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:49:35 2011