
| Sequence ID | dm3.chrX |
|---|---|
| Location | 16,312,220 – 16,312,357 |
| Length | 137 |
| Max. P | 0.829365 |

| Location | 16,312,220 – 16,312,328 |
|---|---|
| Length | 108 |
| Sequences | 14 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 77.47 |
| Shannon entropy | 0.52035 |
| G+C content | 0.49325 |
| Mean single sequence MFE | -32.79 |
| Consensus MFE | -17.35 |
| Energy contribution | -17.34 |
| Covariance contribution | -0.01 |
| Combinations/Pair | 1.52 |
| Mean z-score | -1.07 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.535961 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 16312220 108 - 22422827 CGCAGAUAUUCCAAGAGUACGAGGGCAUGACUGGCCACUCGCUGGAGAAGGCGAUCAAGAAGGAGUUCUCCGGCGAUGUGAUGGAGGGCCUGAUUGCCAUCUACAGGU ......(((((...)))))(((((((.......))).))))(((.(((.((((((((.....(.((((((((.........))))))))))))))))).))).))).. ( -36.00, z-score = -0.59, R) >droSim1.chrX 12557442 108 - 17042790 UGCAGAUAUUCCAAGAGUACGAGGGCAUGACUGGCCACUCGCUGGAGAAGGCGAUCAAGAAGGAGUUCUCCGGCGAUGUGAUGGAGGGCCUGAUUGCCAUCUACAGGU ......(((((...)))))(((((((.......))).))))(((.(((.((((((((.....(.((((((((.........))))))))))))))))).))).))).. ( -36.00, z-score = -0.64, R) >droSec1.super_17 131942 108 - 1527944 UGCAGAUAUUCCAAGAGUACGAGGGCAUGACUGGCCACUCGCUGGAGAAGGCGAUCAAGAAGGAGUUCUCCGGCGACGUGAUGGAGGGCCUGAUUGCCAUCUACAGGU ((.((((.(((((....((((.((.((....)).))..(((((((((((..(..........)..))))))))))))))).)))))(((......))))))).))... ( -36.70, z-score = -0.83, R) >droYak2.chrX 10431515 108 - 21770863 CGCAGAUAUUCCAAGAGUACGAGGGAAUGACUGGCCACUCGCUGGAGAAGGCCAUCAAGAAGGAGUUCUCCGGCGAUGUGAUGGAGGGCCUGAUUGCCAUCUACAAGU ..(((.((((((..(....)...)))))).)))..((((((((((((((..((........))..))))))))))).))).((((.(((......))).))))..... ( -41.50, z-score = -2.68, R) >droEre2.scaffold_4690 7963055 108 + 18748788 UGCAGAUAUUCCAAGAGUACGAGGGCAUGACUGGCCACUCGCUGGAGAAGGCCAUCAAGAAGGAGUUCUCCGGCGAUGUGAUGGAGGGCCUGAUUGCCAUCUACAGGU ((.((((.(((((....(((((((((.......))).)))(((((((((..((........))..)))))))))...))).)))))(((......))))))).))... ( -39.70, z-score = -1.52, R) >droAna3.scaffold_13137 604649 108 - 1373759 CCCAGAUAUUCCAAGAGUACGAGAACAUGACGGGUCACUCGUUGGAGAAGGCCAUCAAAAAGGAGUUUUCUGGUGACAUUAUGGAAGGUCUAAUCGCCAUCUUCAGGU ...((((.(((((.......((((((......((((.(((....)))..)))).((......))))))))((....))...))))).))))....(((.......))) ( -25.10, z-score = 0.56, R) >dp4.chrXL_group1e 9936456 108 - 12523060 UGUAGAUAUUCCAAGAGUAUGAGGGCAUGACCGGCCACUCGCUGGAGAAGGCCAUCAAGAAGGAGUUCUCCGGUGACAUUAUGGAGGGCCUGAUUGCCAUUUUCCGUU .....((((((...))))))...((((...(.((((.((((((((((((..((........))..)))))))))..(.....)))))))).)..)))).......... ( -35.40, z-score = -1.01, R) >droPer1.super_22 1389593 108 + 1688296 UGUAGAUAUUCCAAGAGUAUGAGGGCAUGACUGGCCACUCGCUGGAGAAGGCCAUCAAGAAGGAGUUCUCCGGUGACAUUAUGGAGGGCCUGAUUGCCAUUUUCCGUU .....((((((...))))))...((((.....((((.((((((((((((..((........))..)))))))))..(.....))))))))....)))).......... ( -35.90, z-score = -1.17, R) >droWil1.scaffold_181096 5977389 105 + 12416693 ---AGAUAUUCCAAGAAUAUGAAAACAUGACUGGUCACUCUCUGGAGAAGGCAGUUAAAAAGGAAUUCUCUGGCGAUAUUAUGGAGGGUUUGAUCGCUAUCUACAAGU ---(((.(((((.......((....))((((((.(..(((....)))..).))))))....)))))))).(((((((...((.....))...)))))))......... ( -20.60, z-score = 0.55, R) >droVir3.scaffold_12970 4336865 106 + 11907090 --CAGAUAUUCCAAGAGUACGAAAACAUGACUGGGCACUCGUUCGAGAAGGCGCUCAAAAAGGAGUUCUCGGGCGAUAUUAUGGAGGGUCUAAUUGCCAUCUACAAGU --.((((.(((((..(((.((......)))))......((((((((((....((((......)))))))))))))).....))))).))))................. ( -25.60, z-score = -0.38, R) >droMoj3.scaffold_6482 2151883 106 - 2735782 --CAGAUAUUCCAGGAGUACGAGAGUAUAACUGGACACUCGCUGGAGAAGGCGAUCAAAAAAGAAUUCUCUGGCGAUAUUAUGGAGGGCCUUAUCGCCAUCUACAAGU --.......(((((..((((....))))..)))))...((((..(((((..(..........)..)))))..)))).....((((.(((......))).))))..... ( -37.00, z-score = -3.89, R) >droGri2.scaffold_15203 7041898 108 - 11997470 GAUAGAUAUUCCAAGAGUAUGCGGACAUGACUGGCCAUUCACUGGAGAAAGCCAUCAAAAAGGAAUUCUCUGGCGAUAUUAUGGAAGGUCUAAUUGCUAUCUUCAAGU (((((((((((...))))))........((((..((((((.(..(((((..((........))..)))))..).))....))))..))))......)))))....... ( -29.90, z-score = -1.75, R) >anoGam1.chr2R 17450481 91 + 62725911 -----------------UACGAGAGCCUCGCCGGCCACUCGAUCGAGGAUGCGAUCAAGCGCGAAUUCAGCGGUGCUAUCGAGGAAGGCUUCAAGGCCAUCGGUGAGU -----------------.........((((((((...((((((((......))))).(((((..........)))))...)))...((((....)))).)))))))). ( -32.80, z-score = -0.43, R) >triCas2.ChLG9 11941794 101 - 15222296 -------AUUUCAGGAGUAUCACAGAAUUUCAGGACAUGACAUUGAGAAGGCCAUCAAAAAAGAAUUUUCUGGUGACAUUCAGGAUGGUCUUUUGGCUGUUGUGAGAU -------......(((((........)))))....((..(((.(.(((((((((((.....(((....)))(....)......))))))))))).).)))..)).... ( -26.90, z-score = -1.20, R) >consensus _GCAGAUAUUCCAAGAGUACGAGGGCAUGACUGGCCACUCGCUGGAGAAGGCCAUCAAGAAGGAGUUCUCCGGCGAUAUUAUGGAGGGCCUGAUUGCCAUCUACAGGU ........(((((...................(....)(((((((((((..(..........)..))))))))))).....)))))(((......))).......... (-17.35 = -17.34 + -0.01)



| Location | 16,312,248 – 16,312,357 |
|---|---|
| Length | 109 |
| Sequences | 12 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 70.80 |
| Shannon entropy | 0.64510 |
| G+C content | 0.44297 |
| Mean single sequence MFE | -26.17 |
| Consensus MFE | -15.19 |
| Energy contribution | -14.56 |
| Covariance contribution | -0.63 |
| Combinations/Pair | 1.53 |
| Mean z-score | -0.65 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.829365 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 16312248 109 - 22422827 ---AUUGAAACUCCAACAUAAUCAUACCUAACCGCAGAUAUUCCAAGAGUACGAGGGCAUGACUGGCCACUCGCUGGAGAAGGCGAUCAAGAAGGAGUUCUCCGGCGAUGUG ---..................((((.(((...((....(((((...))))))).))).)))).....((((((((((((((..(..........)..))))))))))).))) ( -25.90, z-score = 0.27, R) >droSim1.chrX 12557470 109 - 17042790 ---AUUGAAACCGCAACUUAAUCCUACCCAACUGCAGAUAUUCCAAGAGUACGAGGGCAUGACUGGCCACUCGCUGGAGAAGGCGAUCAAGAAGGAGUUCUCCGGCGAUGUG ---........(((..(((.(((....((..((.(((.(((((...)))))(((((((.......))).))))))).))..)).))).)))..(((....))).)))..... ( -27.20, z-score = 0.30, R) >droSec1.super_17 131970 109 - 1527944 ---AUUGAAACCGCAACUUAAUCCUACCCAACUGCAGAUAUUCCAAGAGUACGAGGGCAUGACUGGCCACUCGCUGGAGAAGGCGAUCAAGAAGGAGUUCUCCGGCGACGUG ---........(((..(((.(((....((..((.(((.(((((...)))))(((((((.......))).))))))).))..)).))).)))..(((....))).)))..... ( -27.20, z-score = 0.33, R) >droYak2.chrX 10431543 109 - 21770863 ---AUUGAAACGCCAAUUUAAUCCUAACAAUUCGCAGAUAUUCCAAGAGUACGAGGGAAUGACUGGCCACUCGCUGGAGAAGGCCAUCAAGAAGGAGUUCUCCGGCGAUGUG ---........((((..(((.((((....((((...(.....)...))))....)))).))).))))((((((((((((((..((........))..))))))))))).))) ( -35.80, z-score = -2.78, R) >droEre2.scaffold_4690 7963083 108 + 18748788 ---ACUGACACCCC-ACUUAAUGCUACCUUACUGCAGAUAUUCCAAGAGUACGAGGGCAUGACUGGCCACUCGCUGGAGAAGGCCAUCAAGAAGGAGUUCUCCGGCGAUGUG ---.........((-(....(((((..(.((((..............)))).)..)))))...))).((((((((((((((..((........))..))))))))))).))) ( -34.24, z-score = -1.68, R) >droAna3.scaffold_13137 604677 107 - 1373759 -----AAUGAUUCCCAAUAAACGAUCCUAUUCCCCAGAUAUUCCAAGAGUACGAGAACAUGACGGGUCACUCGUUGGAGAAGGCCAUCAAAAAGGAGUUUUCUGGUGACAUU -----.........................((.(((((.(((((....((......)).(((..((((.(((....)))..)))).)))....)))))..))))).)).... ( -21.70, z-score = 0.73, R) >dp4.chrXL_group1e 9936484 104 - 12523060 ------GAUGGAUU-CAAUAGUUCUA-UUCUUUGUAGAUAUUCCAAGAGUAUGAGGGCAUGACCGGCCACUCGCUGGAGAAGGCCAUCAAGAAGGAGUUCUCCGGUGACAUU ------((((....-......(((((-(((((.(........).))))))).)))(((.......)))..(((((((((((..((........))..))))))))))))))) ( -30.20, z-score = -0.85, R) >droPer1.super_22 1389621 105 + 1688296 ------GAUGGAUCACAAUAGUUCUA-UUCUUUGUAGAUAUUCCAAGAGUAUGAGGGCAUGACUGGCCACUCGCUGGAGAAGGCCAUCAAGAAGGAGUUCUCCGGUGACAUU ------((((...........(((((-(((((.(........).))))))).)))(((.......)))..(((((((((((..((........))..))))))))))))))) ( -30.20, z-score = -0.76, R) >droWil1.scaffold_181096 5977417 102 + 12416693 ---------AUUCGAAUAAACUGAUUGCUCUU-UUAGAUAUUCCAAGAAUAUGAAAACAUGACUGGUCACUCUCUGGAGAAGGCAGUUAAAAAGGAAUUCUCUGGCGAUAUU ---------..............((((((...-..(((.(((((.......((....))((((((.(..(((....)))..).))))))....))))))))..))))))... ( -18.70, z-score = 0.12, R) >droVir3.scaffold_12970 4336893 111 + 11907090 AACAAUAUUAUUUAAA-AAUCUAUUUGAACUUACCAGAUAUUCCAAGAGUACGAAAACAUGACUGGGCACUCGUUCGAGAAGGCGCUCAAAAAGGAGUUCUCGGGCGAUAUU ..........((((((-......))))))....((((.(((((...))))).(....)....))))....((((((((((....((((......)))))))))))))).... ( -21.50, z-score = -0.56, R) >droMoj3.scaffold_6482 2151911 103 - 2735782 ---------AUCAACAUUCGAUGCACACUCUUUUCAGAUAUUCCAGGAGUACGAGAGUAUAACUGGACACUCGCUGGAGAAGGCGAUCAAAAAAGAAUUCUCUGGCGAUAUU ---------................................(((((..((((....))))..)))))...((((..(((((..(..........)..)))))..)))).... ( -30.10, z-score = -2.98, R) >apiMel3.Group11 2027816 96 - 12576330 --------------AAUGUACCACAACUGAA--ACAAAUAUUUGAAGAAUAUGAAAAUAUUACUGGUAACAACAUUGAAACUGCAAUUAAAAACGAGUUUUCGGGUGAUAUU --------------.......(((..(((((--(.............(((((....)))))(((.((......((((......)))).....)).))))))))))))..... ( -11.30, z-score = 0.08, R) >consensus _____UGAAACCCCAACUUAAUCCUACCUAUUUGCAGAUAUUCCAAGAGUACGAGGGCAUGACUGGCCACUCGCUGGAGAAGGCCAUCAAGAAGGAGUUCUCCGGCGAUAUU ......................................(((((...)))))...................(((((((((((..(..........)..))))))))))).... (-15.19 = -14.56 + -0.63)


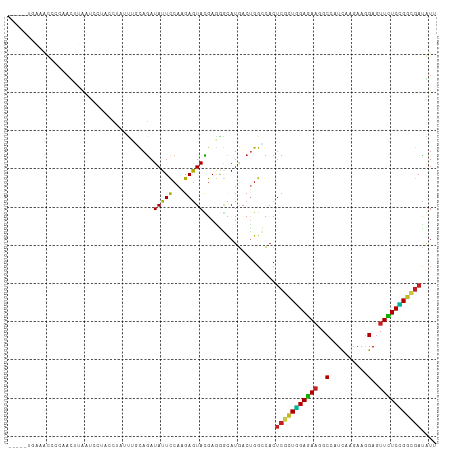
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:49:31 2011