
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 11,508,911 – 11,509,011 |
| Length | 100 |
| Max. P | 0.993051 |

| Location | 11,508,911 – 11,509,011 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 95.60 |
| Shannon entropy | 0.07493 |
| G+C content | 0.33200 |
| Mean single sequence MFE | -21.78 |
| Consensus MFE | -18.10 |
| Energy contribution | -19.10 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.83 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.772049 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
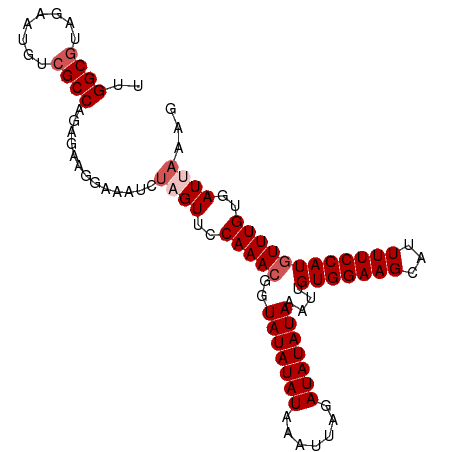
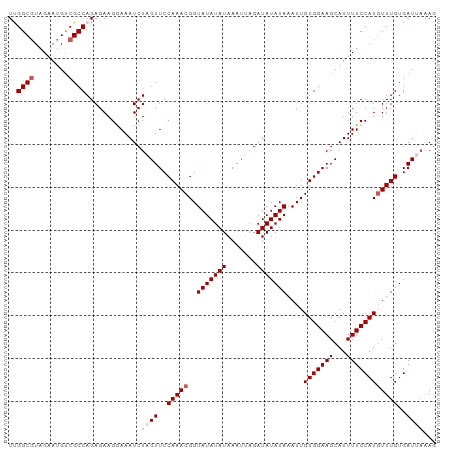
>dm3.chr2L 11508911 100 + 23011544 UUGGCUUAGAAUGUCGCCAGAGAAGGAAAUCUAGUUCCAAACGGUAUAUAUAAAUUAGAUAUAUAAAUUGUGGAAGCAUUUUCCAUUUUUGUGAUUAAAG ............(((((.(((((.(((((.....(((((.....(((((((.......))))))).....)))))....))))).))))))))))..... ( -21.30, z-score = -1.90, R) >droSim1.chr2L 11324801 100 + 22036055 UUGGCGUAGAAUGUCGCCAGAGAAGGAAAUCUAGUUCCAAACGGUAUAUAUAAAUUAGAUAUAUAAAUUGUGGAAGCAUUUUCCAUGUUUGUGAUUAAAG ((((((........))))))...........((((..(((((..(((((((.......)))))))....(((((((...))))))))))))..))))... ( -23.10, z-score = -2.58, R) >droYak2.chr2L 7915399 100 + 22324452 UCGGCGUAGAAUGCCGCCAGAGAAGAAAAUCUAGUUCCAAACGGUAUAUAUAAAUAAGAUAUAUAAAUUGUGGAAGCAUUUUCCAUGUUUGUGAUAAAAG ..((((........))))(((........))).((..(((((..(((((((.......)))))))....(((((((...))))))))))))..))..... ( -20.30, z-score = -2.01, R) >droEre2.scaffold_4929 12717782 100 - 26641161 UCGGCGUAGAAUGUCGCCAGAGAAGAAAAUCUAGUUCCAAACGGUAUAUAUAAAUAAGAUAUAUAAAUUGUGGAAGCAUUUUCCAUGUUUGUGAUACGAG ..((((........))))(((........))).(((((((((..(((((((.......)))))))....(((((((...)))))))))))).)).))... ( -21.10, z-score = -1.73, R) >droSec1.super_3 6906027 100 + 7220098 UUGGCGUAGAAUGUCGCCAGAGAAGGAAAUCUAGUUCCAAACGGUAUAUAUAAAUUAGAUAUAUAAAUUGUGGAAGCAUUUUCCAUGUUUGUGAUUAAAG ((((((........))))))...........((((..(((((..(((((((.......)))))))....(((((((...))))))))))))..))))... ( -23.10, z-score = -2.58, R) >consensus UUGGCGUAGAAUGUCGCCAGAGAAGGAAAUCUAGUUCCAAACGGUAUAUAUAAAUUAGAUAUAUAAAUUGUGGAAGCAUUUUCCAUGUUUGUGAUUAAAG ..((((........)))).............((((..(((((..(((((((.......)))))))....(((((((...))))))))))))..))))... (-18.10 = -19.10 + 1.00)

| Location | 11,508,911 – 11,509,011 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 95.60 |
| Shannon entropy | 0.07493 |
| G+C content | 0.33200 |
| Mean single sequence MFE | -18.66 |
| Consensus MFE | -17.10 |
| Energy contribution | -16.82 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.05 |
| Structure conservation index | 0.92 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.58 |
| SVM RNA-class probability | 0.993051 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
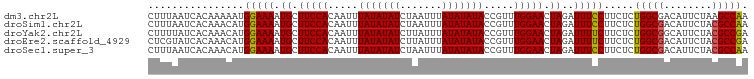
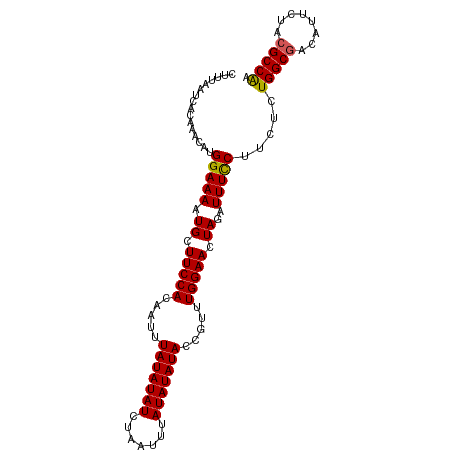
>dm3.chr2L 11508911 100 - 23011544 CUUUAAUCACAAAAAUGGAAAAUGCUUCCACAAUUUAUAUAUCUAAUUUAUAUAUACCGUUUGGAACUAGAUUUCCUUCUCUGGCGACAUUCUAAGCCAA ................(((((.((.(((((.....(((((((.......))))))).....))))).))..))))).....((((..........)))). ( -16.50, z-score = -2.17, R) >droSim1.chr2L 11324801 100 - 22036055 CUUUAAUCACAAACAUGGAAAAUGCUUCCACAAUUUAUAUAUCUAAUUUAUAUAUACCGUUUGGAACUAGAUUUCCUUCUCUGGCGACAUUCUACGCCAA ................(((((.((.(((((.....(((((((.......))))))).....))))).))..))))).....(((((........))))). ( -18.70, z-score = -3.52, R) >droYak2.chr2L 7915399 100 - 22324452 CUUUUAUCACAAACAUGGAAAAUGCUUCCACAAUUUAUAUAUCUUAUUUAUAUAUACCGUUUGGAACUAGAUUUUCUUCUCUGGCGGCAUUCUACGCCGA ......((.(((((.(((((.....))))).....((((((.......))))))....))))))).(((((........)))))((((.......)))). ( -20.70, z-score = -3.51, R) >droEre2.scaffold_4929 12717782 100 + 26641161 CUCGUAUCACAAACAUGGAAAAUGCUUCCACAAUUUAUAUAUCUUAUUUAUAUAUACCGUUUGGAACUAGAUUUUCUUCUCUGGCGACAUUCUACGCCGA ((.((.((.(((((.(((((.....))))).....((((((.......))))))....))))))))).))...........(((((........))))). ( -18.70, z-score = -2.55, R) >droSec1.super_3 6906027 100 - 7220098 CUUUAAUCACAAACAUGGAAAAUGCUUCCACAAUUUAUAUAUCUAAUUUAUAUAUACCGUUUGGAACUAGAUUUCCUUCUCUGGCGACAUUCUACGCCAA ................(((((.((.(((((.....(((((((.......))))))).....))))).))..))))).....(((((........))))). ( -18.70, z-score = -3.52, R) >consensus CUUUAAUCACAAACAUGGAAAAUGCUUCCACAAUUUAUAUAUCUAAUUUAUAUAUACCGUUUGGAACUAGAUUUCCUUCUCUGGCGACAUUCUACGCCAA ................(((((.((.(((((.....(((((((.......))))))).....))))).))..))))).....(((((........))))). (-17.10 = -16.82 + -0.28)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:32:56 2011