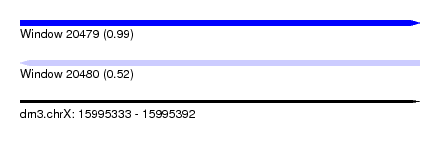

| Sequence ID | dm3.chrX |
|---|---|
| Location | 15,995,333 – 15,995,392 |
| Length | 59 |
| Max. P | 0.987875 |

| Location | 15,995,333 – 15,995,392 |
|---|---|
| Length | 59 |
| Sequences | 5 |
| Columns | 60 |
| Reading direction | forward |
| Mean pairwise identity | 94.44 |
| Shannon entropy | 0.09919 |
| G+C content | 0.33452 |
| Mean single sequence MFE | -14.74 |
| Consensus MFE | -13.72 |
| Energy contribution | -13.24 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.93 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.30 |
| SVM RNA-class probability | 0.987875 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
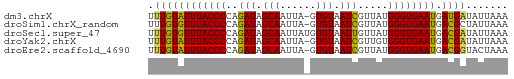
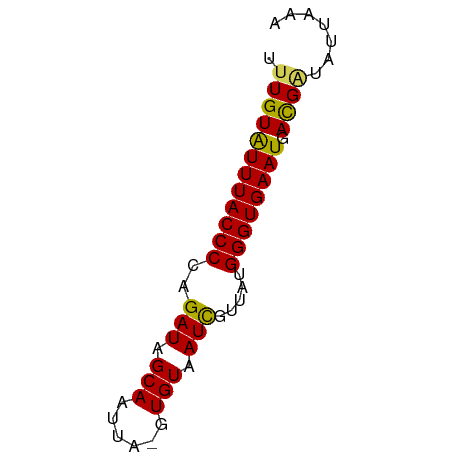
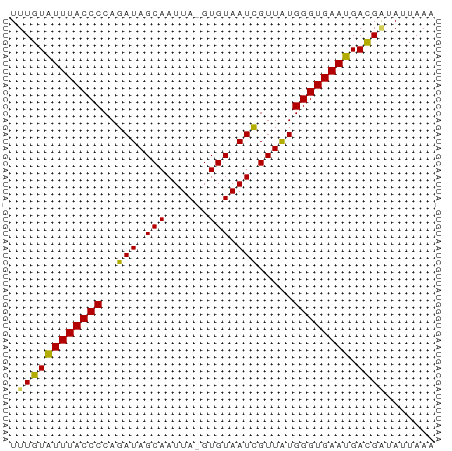
>dm3.chrX 15995333 59 + 22422827 UUUGUAUUUACCCCAGAUAGCAAUUA-GUGUAAUCGUUAUGGGUGAAUGAUGAUAUUAAA ....(((((((((..(((.(((....-.))).))).....)))))))))........... ( -13.10, z-score = -2.30, R) >droSim1.chrX_random 4129404 59 + 5698898 UUUGUGUUUACCCCAGAUAGCAAUUA-GUGUAAUCGUUAUGGGUGAAUGACGCUAUUAAA ...((((((((((..(((.(((....-.))).))).....)))))))).))......... ( -13.00, z-score = -1.74, R) >droSec1.super_47 53345 60 + 216421 UUUGUGUUUACCCCAGAUAGCAAUUAUGUGUAAUUGUUAUGGGUGAAUGACGAUAUUAAA .((((((((((((...((((((((((....)))))))))))))))))).))))....... ( -18.90, z-score = -4.55, R) >droYak2.chrX 10143020 59 + 21770863 UUUGUAUUUACCCCAGAUAGCAAUUA-GUGUAAUCGUUGUGGGUGAAUGACGAUAUUAAA .(((((((((((((((((.(((....-.))).)))..)).)))))))).))))....... ( -15.20, z-score = -3.00, R) >droEre2.scaffold_4690 7665080 59 - 18748788 UUUGUAUUUACCCCAGAUAGCAAUUA-GUGUAAUCGUUAUGGGUGAAUGACGGUACUAAA .((((((((((((..(((.(((....-.))).))).....)))))))).))))....... ( -13.50, z-score = -1.54, R) >consensus UUUGUAUUUACCCCAGAUAGCAAUUA_GUGUAAUCGUUAUGGGUGAAUGACGAUAUUAAA .((((((((((((...(((((.((((....)))).))))))))))))).))))....... (-13.72 = -13.24 + -0.48)

| Location | 15,995,333 – 15,995,392 |
|---|---|
| Length | 59 |
| Sequences | 5 |
| Columns | 60 |
| Reading direction | reverse |
| Mean pairwise identity | 94.44 |
| Shannon entropy | 0.09919 |
| G+C content | 0.33452 |
| Mean single sequence MFE | -7.62 |
| Consensus MFE | -7.14 |
| Energy contribution | -6.98 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.06 |
| Structure conservation index | 0.94 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.517210 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
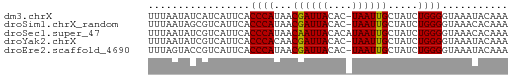
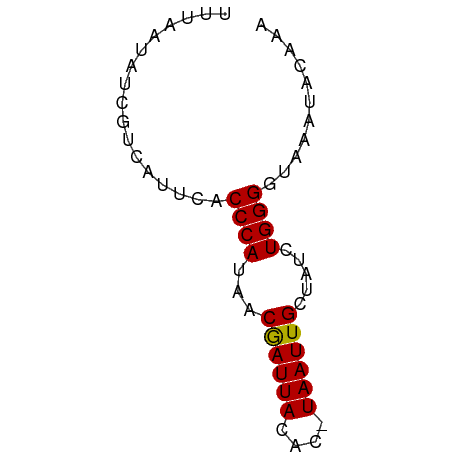
>dm3.chrX 15995333 59 - 22422827 UUUAAUAUCAUCAUUCACCCAUAACGAUUACAC-UAAUUGCUAUCUGGGGUAAAUACAAA ............(((.(((((((.((((((...-)))))).)))...)))).)))..... ( -7.00, z-score = -1.22, R) >droSim1.chrX_random 4129404 59 - 5698898 UUUAAUAGCGUCAUUCACCCAUAACGAUUACAC-UAAUUGCUAUCUGGGGUAAACACAAA .........((......((((((.((((((...-)))))).))..))))......))... ( -6.80, z-score = -0.42, R) >droSec1.super_47 53345 60 - 216421 UUUAAUAUCGUCAUUCACCCAUAACAAUUACACAUAAUUGCUAUCUGGGGUAAACACAAA .........((......((((((.((((((....)))))).))..))))......))... ( -7.80, z-score = -1.66, R) >droYak2.chrX 10143020 59 - 21770863 UUUAAUAUCGUCAUUCACCCACAACGAUUACAC-UAAUUGCUAUCUGGGGUAAAUACAAA .........((.(((.((((.((..(((..((.-....))..))))))))).)))))... ( -8.30, z-score = -1.54, R) >droEre2.scaffold_4690 7665080 59 + 18748788 UUUAGUACCGUCAUUCACCCAUAACGAUUACAC-UAAUUGCUAUCUGGGGUAAAUACAAA .........((.(((.(((((((.((((((...-)))))).)))...)))).)))))... ( -8.20, z-score = -0.47, R) >consensus UUUAAUAUCGUCAUUCACCCAUAACGAUUACAC_UAAUUGCUAUCUGGGGUAAAUACAAA .................((((...((((((....)))))).....))))........... ( -7.14 = -6.98 + -0.16)
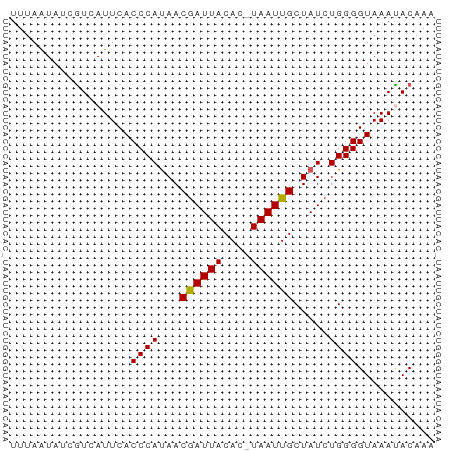
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:48:28 2011