| Sequence ID | dm3.chrX |
|---|---|
| Location | 15,988,205 – 15,988,296 |
| Length | 91 |
| Max. P | 0.596677 |

| Location | 15,988,205 – 15,988,296 |
|---|---|
| Length | 91 |
| Sequences | 8 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 63.85 |
| Shannon entropy | 0.63548 |
| G+C content | 0.56301 |
| Mean single sequence MFE | -31.99 |
| Consensus MFE | -11.46 |
| Energy contribution | -12.65 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.596677 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
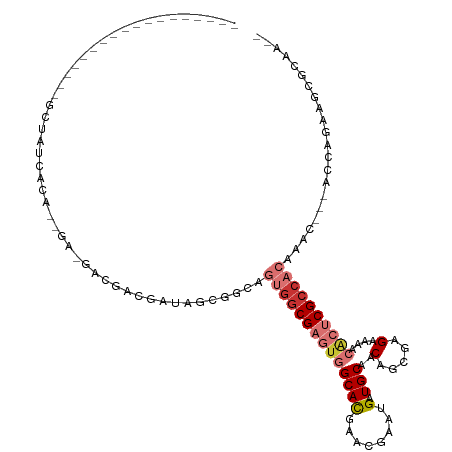
>dm3.chrX 15988205 91 - 22422827 ------------------GCUAUCCUA------AUAACGAUAGCGGUAGUGGCGAGUGGCAUGAACGAAUGAUGCUACAGCGAGAAAACGCUCGCCACAAAC---AUCAGAAGCGGAA-- ------------------((((((...------.....))))))....((((((((((((((.(.....).))))))).(((......))))))))))....---.............-- ( -32.60, z-score = -3.97, R) >droPer1.super_22 1068656 120 + 1688296 GCCAUACUAGCGGCGAGCGGGACGAGAGCGAUGGCGAUGGCAGCGGCAGUGGCGAAUGGCACGGGAAAUUGAUGCAGCAUCGCGAGCAGAGUCGCCACAGACACGACCAGAAGCGCAAGG (((........))).............(((.(((((.((((....)).(((((((...((.((.((..(((...)))..)).)).))....)))))))...)))).)))....))).... ( -36.70, z-score = 0.53, R) >dp4.chrXL_group1e 9589490 120 - 12523060 GCCAUACUAGCGGCGAGCGGGACGAGAGCGAUGGCGAUGGCAGCGGCAGUGGCGAAUGGCACGGGAAAUUGAUGCAGCAUCGCGAGCAGAGUCGCCACAGACACGACCAGAAGCGCAAGG (((........))).............(((.(((((.((((....)).(((((((...((.((.((..(((...)))..)).)).))....)))))))...)))).)))....))).... ( -36.70, z-score = 0.53, R) >droSim1.chrX_random 4122286 91 - 5698898 ------------------GCUAUCCCA------AUCACGAUAGCGGUAGUGGCGAGUGGCAUGAACGAAUGAUGCUACAGCGAGAAAACACUCGCCACAAAC---AUCAGAAGCGGAA-- ------------------((((((...------.....))))))....((((((((((.(((......))).(((....)))......))))))))))....---.............-- ( -30.60, z-score = -3.44, R) >droSec1.super_47 46297 91 - 216421 ------------------GCUAUCCCA------AUCACGAUAGCGGUAGUGGCGAGUGGCAUGAACGAAUGAUGCUACAGCGAGAAAACACUCGCCACAAAC---AUCAGAAGCGGAA-- ------------------((((((...------.....))))))....((((((((((.(((......))).(((....)))......))))))))))....---.............-- ( -30.60, z-score = -3.44, R) >droEre2.scaffold_4690 7658324 97 + 18748788 ------------------GCAAUCCUAAUGACGAUAACGAUAGCGGCAGUGGCGAGUGGCAUGAACGAAUGAUGCUACAGCGUGAAAACGCUCGCCACAAAC---ACCAGAAGCGCAA-- ------------------((.................((....))...((((((((((((((.(.....).))))))).((((....)))))))))))....---.........))..-- ( -31.50, z-score = -2.59, R) >droMoj3.scaffold_6308 2959650 97 - 3356042 ------------------AGCGCCAUGAUGAUGACGAUGACAGCGCCAGCGGCGAGUGGCACGAGCGCAUGAUGCAACAUCGUGAGCACGCACGCCAGCGUCACGAGAAGAAGCG----- ------------------.((..(((....))).((.((((.(((((...)))...((((.((.((.(((((((...))))))).)).))...)))))))))))).......)).----- ( -30.60, z-score = 0.73, R) >droGri2.scaffold_15203 4454377 97 - 11997470 ------------------GUCAUCACCGGCAGGACGAUGAUAGUGGCAGCGGCGAGUGGCACGAACGGAGGAUGCAACAUCGUGAGCACUCACGAACGCAGCACGAACAGAGACG----- ------------------(((((((.((......)).)))).(((.(.(((..(((((.(((((...(......)....)))))..))))).....))).)))).......))).----- ( -26.60, z-score = 0.06, R) >consensus __________________GCUAUCACA__GA_GACGACGAUAGCGGCAGUGGCGAGUGGCACGAACGAAUGAUGCAACAGCGAGAAAACACUCGCCACAAAC___ACCAGAAGCGCAA__ ................................................((((((((((((((........).)))..(.....)....))))))))))...................... (-11.46 = -12.65 + 1.19)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:48:25 2011