
| Sequence ID | dm3.chrX |
|---|---|
| Location | 15,980,689 – 15,980,792 |
| Length | 103 |
| Max. P | 0.545072 |

| Location | 15,980,689 – 15,980,792 |
|---|---|
| Length | 103 |
| Sequences | 7 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 72.45 |
| Shannon entropy | 0.48268 |
| G+C content | 0.41732 |
| Mean single sequence MFE | -28.34 |
| Consensus MFE | -15.76 |
| Energy contribution | -14.13 |
| Covariance contribution | -1.62 |
| Combinations/Pair | 1.46 |
| Mean z-score | -1.15 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.545072 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 15980689 103 - 22422827 CCUGCUAAAGGCAAAUGUCAAACCUUCGUAACGGUUAUGCCUUUCAUAGCAUGUGGCCUUCAAUUGGUUUAUAAAAUGCAAAAAGCAGGUUAUGUACA-UGCAA----------- ......(((((((.......((((........)))).)))))))....((((((((((.......))).(((((..(((.....)))..)))))))))-)))..----------- ( -25.30, z-score = -1.03, R) >droSim1.chrX_random 4114393 103 - 5698898 CCUGCUAAAGGCAAAUGUCAAACCUUCGUAACGGUUAUGCCUUUCAUAGCAUGUGGCCUUCAAUUGGUUUAUAAAAUGCAAAAAGCAGGUUAUGUACA-UGCAA----------- ......(((((((.......((((........)))).)))))))....((((((((((.......))).(((((..(((.....)))..)))))))))-)))..----------- ( -25.30, z-score = -1.03, R) >droSec1.super_47 38779 103 - 216421 CCUGCUAAAGGCAAAUGUCAAACCUUCGUAACGGUUAUGCCUUUCAUAGCAUGUGGCCUUCAAUUGGUUUAUAAAAUGCAAAAAGCAGGUUAUGUACA-UGCAA----------- ......(((((((.......((((........)))).)))))))....((((((((((.......))).(((((..(((.....)))..)))))))))-)))..----------- ( -25.30, z-score = -1.03, R) >droYak2.chrX 10129191 103 - 21770863 GCUGCUGAAGGCAAGUGCCAAACCUUCGUAACGGUUAUGCCUUUCAUAGCAUGUGGCCUUCAAUUGGUUUAUAAAAUGCAAAAAGCAGGUUAUGUACA-UGCAA----------- ..((((((((((..((((..((((........))))(((.....))).))))...)))))))....((.(((((..(((.....)))..))))).)).-.))).----------- ( -28.40, z-score = -0.72, R) >droEre2.scaffold_4690 7651082 103 + 18748788 CCCUCUGAAGGCAAAUGCCAAACCUUCGUAACGGCUAUGCCUUGUAUGGCAUUUGGCCUUCAAUUGGUUUAUAAAAUGCAAAAAGCAGGUUAUGUACA-UGCAU----------- .....(((((((((((((((.((...((((.....))))....)).)))))))).)))))))....((.(((((..(((.....)))..))))).)).-.....----------- ( -31.10, z-score = -2.00, R) >dp4.chrXL_group1e 9580729 110 - 12523060 --CUCAAACUUCCAGUGGCGGGGCUU---GAGGCUUGGCGAUUUCAAUGUGUGUGGCCUUCAGAUGGUUUAAAAAAGACACACAGCCAGUAGCAUAUACUACGAGUAUAGGAUAU --(((((.(((((....).)))).))---)))((((((((....)..(((((((((((.......))))........)))))))))))..))).(((((.....)))))...... ( -27.90, z-score = -0.06, R) >droPer1.super_22 1059804 105 + 1688296 --CUCAAGCUUCCAGUGGCGGGGCUU---GAGGCUUGGCGAUUUCAAUGUGUGUGGCCUUCAGAUGGUUUAAAAAAGACACACAGCCAGUAGUAUAUACUACGAGUAUGA----- --(((((((((((....).)))))))---)))((((((((....)..(((((((((((.......))))........)))))))))).(((((....)))))))))....----- ( -35.10, z-score = -2.21, R) >consensus CCUGCUAAAGGCAAAUGUCAAACCUUCGUAACGGUUAUGCCUUUCAUAGCAUGUGGCCUUCAAUUGGUUUAUAAAAUGCAAAAAGCAGGUUAUGUACA_UGCAA___________ ......(((((((...(((.............)))..)))))))....((((((((((.......))).(((((..(((.....)))..))))))))).)))............. (-15.76 = -14.13 + -1.62)

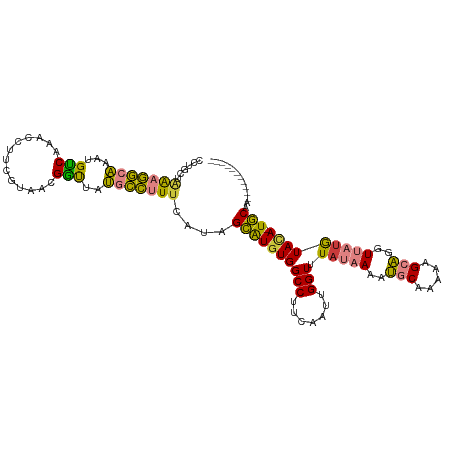

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:48:23 2011