| Sequence ID | dm3.chrX |
|---|---|
| Location | 15,957,547 – 15,957,690 |
| Length | 143 |
| Max. P | 0.981067 |

| Location | 15,957,547 – 15,957,690 |
|---|---|
| Length | 143 |
| Sequences | 7 |
| Columns | 144 |
| Reading direction | forward |
| Mean pairwise identity | 68.85 |
| Shannon entropy | 0.59033 |
| G+C content | 0.48753 |
| Mean single sequence MFE | -45.33 |
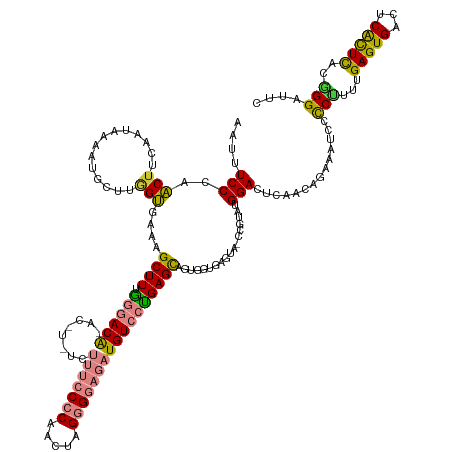
| Consensus MFE | -20.37 |
| Energy contribution | -20.38 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.39 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.06 |
| SVM RNA-class probability | 0.981067 |
| Prediction | RNA |
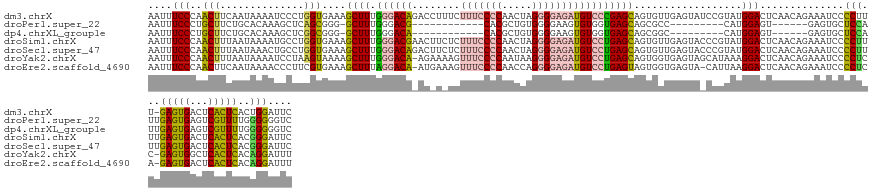
Download alignment: ClustalW | MAF
>dm3.chrX 15957547 143 + 22422827 AAUUUCCCAACUUCAAUAAAAUCCCUGGUGAAAGCUUUGGGACAGACCUUUCUUUCCCCAACUAGGGGAGAUGUCCCGAGCAGUGUUGAGUAUCCGUAUGGACUCAACAGAAAUCCCCUUU-GAGUGACUCACUCACUGGAUUC ..(((((((................))).))))((((.((((((((....))(((((((.....)))))))))))))))))..(((((((..(((....))))))))))..(((((....(-(((((...))))))..))))). ( -46.79, z-score = -2.75, R) >droPer1.super_22 1022849 116 - 1688296 AAUUUCCCUGCUUCUGCACAAAGCUCAGCGGG-GCUUUGGGACG------------CACGCUGUGGGGAAGUGUGGUGAGCAGCGCC---------CAUGGAGU------GAGUGCUCCAUUGAGUGAGUCGUUUUGGGGGGUC .((..(((.((..((((.((((((((....))-))))))..((.------------((((((.......))))))))..))))((((---------.(((((((------....))))))).).)))....))...)))..)). ( -45.10, z-score = -0.74, R) >dp4.chrXL_group1e 9545917 116 + 12523060 AAUUUCCCUGCUUCUGCACAAAGCUCGGCGGG-GCUUUGGGACA------------CACGCUGUGGGGAAGUGUGGUGAGCAGCGGC---------CAUGGAGU------GAGUGCUCCAUUGAGUGAGUCGUUUUGGGGGGUC .((..(((.(((.((((.((((((((....))-))))))..((.------------((((((.......))))))))..)))).)))---------.(((((((------....)))))))...............)))..)). ( -45.50, z-score = -1.14, R) >droSim1.chrX 12318274 144 + 17042790 AAUUUCCCAACUUUAAUAAAAUGCCUGGUGAAAGCUUUGGGACGAACUUCUCUUUCCCCAACUAGGGGAGAUGUCCUGAGCAGUGUUGAGUACCCGUAUGGACUCAACAGAAAUCCCCUUUUGAGUGACUCACUCACGGGAUUC ..(((((((................))).))))((.(..((((((.....))(((((((.....))))))).))))..)))..((((((((.((.....))))))))))..((((((....((((((...)))))).)))))). ( -48.09, z-score = -2.94, R) >droSec1.super_47 10511 144 + 216421 AAUUUCCCAACUUUAAUAAACUGCCUGGUGAAAGCUUUGGGACAGACUUCUCUUUCCCCAACUAGGGGAGAUGUCCUGAGCAGUGUUGAGUACCCGUAUGGACUCAACAGAAAUCCCCUUUUGAGUGACUCACUCACGGGAUUC ...................(((((..(((....)))(..(((((((....))(((((((.....))))))))))))..))))))(((((((.((.....)))))))))...((((((....((((((...)))))).)))))). ( -49.80, z-score = -3.14, R) >droYak2.chrX 10106132 142 + 21770863 AAUUUCCCAACUUUAAUAAAAUCCUAAGUAAAAGCUUUGGGACA-AGAAAAGUUUCCCCAAUAAGGGGAGAUGUCCUGAGCAGUGGUGAGUAGCAUAAAGGACUCAACAGAAAUCCCCUCC-GAGUGGCUCACUCACAGGAUUU ..................(((((((........((.(..(((((-.......(((((((.....))))))))))))..))).(((((((((.((....(((...............)))..-..)).)))))).)))))))))) ( -42.77, z-score = -2.49, R) >droEre2.scaffold_4690 7628227 141 - 18748788 AAUUUCCCAACUUCAAUAAAACCCUUCGUGAAAGCUUUAGGACA-AUGAAAGUUUCCCCAACCAGGGGAGAUGUCCUGAGUAGUGGUGAGUA-CAUUAAGGACUCAACAGAAAUCCCCUCA-GAGUGACUCACUCACAGGAUUU ....(((....................((....))(((((((((-.......(((((((.....))))))))))))))))..(((((((((.-((((.(((...............)))..-.)))))))))).))).)))... ( -39.27, z-score = -1.51, R) >consensus AAUUUCCCAACUUCAAUAAAAUGCUUGGUGAAAGCUUUGGGACA_AC_U_UCUUUCCCCAACUAGGGGAGAUGUCCUGAGCAGUGGUGAGUA_CCGUAUGGACUCAACAGAAAUCCCCUUUUGAGUGACUCACUCACGGGAUUC ....(((..(((..............)))....((((.((((((........(((((((.....)))))))))))))))))..................)))..............(((...(((((...)))))..))).... (-20.37 = -20.38 + 0.01)
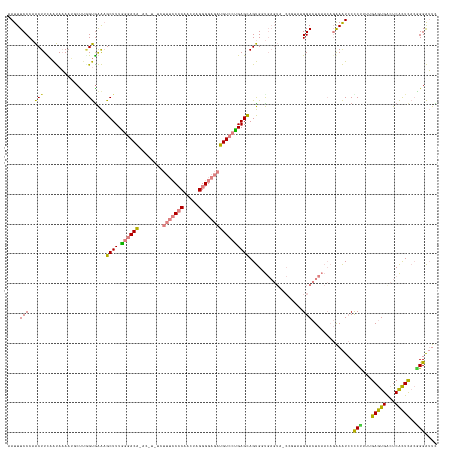
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:48:15 2011