| Sequence ID | dm3.chrX |
|---|---|
| Location | 15,804,875 – 15,805,108 |
| Length | 233 |
| Max. P | 0.879832 |

| Location | 15,804,875 – 15,804,993 |
|---|---|
| Length | 118 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.33 |
| Shannon entropy | 0.26043 |
| G+C content | 0.44613 |
| Mean single sequence MFE | -33.00 |
| Consensus MFE | -21.79 |
| Energy contribution | -23.57 |
| Covariance contribution | 1.78 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.66 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.554538 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 15804875 118 - 22422827 UAACGAAUAUAAAAUUUCAGGGGAUUACAGCAUCCCUCUGACG-GAAAGCGAAGUCGGCGGACGCGAGUGUUCCACU-CAACUCUCGUGUGCUGUUUCAAAGUUUUCUCUGAAUCCUUAA ...............(((((((((..(((((...((.(((((.-.........))))).))(((((((.(((.....-.))).)))))))))))).........)))))))))....... ( -35.70, z-score = -2.32, R) >droSim1.chrX 12223988 118 - 17042790 UAACGAAUAUAAAAUUUCAGGGGAUUACAGCAUCCCUCUGACG-GAAAGCGAAGUCGGCGGACGCGAGUGUUCCACU-CAACUCUCGUGUGCUGUUUCAAAGUUUUCUCUGAAUCCUUAA ...............(((((((((..(((((...((.(((((.-.........))))).))(((((((.(((.....-.))).)))))))))))).........)))))))))....... ( -35.70, z-score = -2.32, R) >droSec1.super_37 381312 118 - 454039 UAACGAAUAUAAAAUUUCAGGGGAUUACAGCAUCCCUCUGACG-GAAAGCGAAGUCGGCGGACGCGAGUGUUCCACU-CAACUCUCGUGUGCUGUUUCAAAGUUUUCUCUGAAUCCUUAA ...............(((((((((..(((((...((.(((((.-.........))))).))(((((((.(((.....-.))).)))))))))))).........)))))))))....... ( -35.70, z-score = -2.32, R) >droYak2.chrX 9995709 118 - 21770863 UAACGAAUAUAAAAUUUCAGGGGAUUACAGCAUCCCUCUGACGCGAAAGCGAAGUCGGCGGACGCGAGUGUUCCACU-CAACUCUCGUGCGCUG-UUAAAAGUUUUCUCCGAAUCCUUAA ((((((..........(((((((((......))))).))))(((....)))...))((((.(((.((((........-..)))).))).)))))-))).(((..(((...)))..))).. ( -37.10, z-score = -2.60, R) >droEre2.scaffold_4690 7505565 117 + 18748788 UAACGAAUAUAAAAUUUCAGGGGAUUACAGCAUCCCUCUGACG-GUAAGCGAAGUCGGCGGACGCGAGUGUUCCACU-CAACUCUCGUGUGCUG-UUAAAAGUUUUCUCUGAAUCCUUAA ...............(((((((((..(((((...((.(((((.-.........))))).))(((((((.(((.....-.))).)))))))))))-)........)))))))))....... ( -35.80, z-score = -2.61, R) >droPer1.super_66 57063 117 + 382987 UGACAAAUGUAAGGUAUCUCGAGAUUACAGCACCGCUCAGCUC-AAAAGCCCAGACCGUGAAAG-AAACGUUUCACUAGAAGUCCCCUUUGAUA-UUAAAAGUUUCCUCUGUACACUUGA ..................(((((..((((((((.(((..(((.-...)))..)).).)))...(-((((.........((((....))))....-......)))))..)))))..))))) ( -18.01, z-score = 0.83, R) >consensus UAACGAAUAUAAAAUUUCAGGGGAUUACAGCAUCCCUCUGACG_GAAAGCGAAGUCGGCGGACGCGAGUGUUCCACU_CAACUCUCGUGUGCUG_UUAAAAGUUUUCUCUGAAUCCUUAA ................(((((((((......))))).))))...((((((.....(((((.(((.((((...........)))).))).))))).......))))))............. (-21.79 = -23.57 + 1.78)


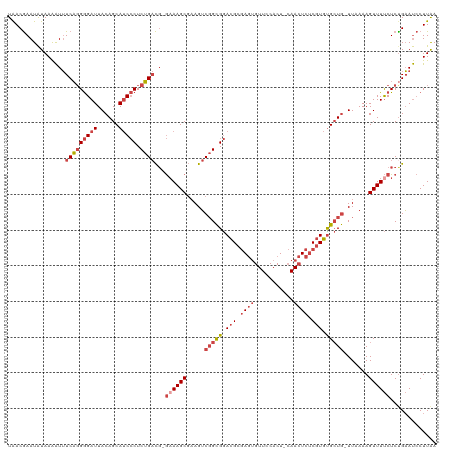
| Location | 15,804,993 – 15,805,108 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 74.37 |
| Shannon entropy | 0.50153 |
| G+C content | 0.39913 |
| Mean single sequence MFE | -30.47 |
| Consensus MFE | -14.34 |
| Energy contribution | -14.40 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.48 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.879832 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 15804993 115 - 22422827 AGUGGCGGGCGAGAACAAUUGAAUCAGCUAAUUGUUCCACA---CACAUUUGUGUGGGUAAUUUACGUGAAUGAAAAAUUAUUUUCGUACAGCACGUCUCUUGUCUUCUUCGGGAAUA .(.((.(((((((((((((((.......)))))))((((((---((....))))))))......(((((.(((((((....)))))))....))))).)))))))).)).)....... ( -32.00, z-score = -2.11, R) >droSim1.chrX 12224106 115 - 17042790 AGUGGCGGGCGAGAACAAUUGAAUCAGCUAAUUGUUCCACA---CACAUUUGUGUGGGUAAUUUACGUGAAUGAAAAAUUAUUUUCGUACAGCACGUCUCUUGUCUUCUUCGGGAAUA .(.((.(((((((((((((((.......)))))))((((((---((....))))))))......(((((.(((((((....)))))))....))))).)))))))).)).)....... ( -32.00, z-score = -2.11, R) >droSec1.super_37 381430 114 - 454039 AGUGGCGGGCGAGAACAAUUGAAUCAGCUAAUUGUUCCGCA---CACAUUUGUGGUGGGUAUUUACGUGAAUGAAAAAUGAUUUUCGUACAGCACGUCUC-UGUCUUCUUCGGGAAUA ...((((((((.(((((((((.......)))))))))))).---(((....)))..........(((((.(((((((....)))))))....)))))..)-))))(((....)))... ( -31.30, z-score = -1.12, R) >droYak2.chrX 9995827 115 - 21770863 AGCAAACGACAAGAACAAUUGAAUCAGCUAAUUGUUCCGCA---CACAUUUGUGUGGGUAAUUUACGUGAAUGAAAAAUUAUUUUCGUACAGCACGUCUCUUGUCUUCUUCGGGAAUA .......((((((((((((((.......)))))))((((((---((....))))))))......(((((.(((((((....)))))))....))))).)))))))(((....)))... ( -32.50, z-score = -3.16, R) >droEre2.scaffold_4690 7505682 114 + 18748788 AUUUCAAGACAAGAACAAUUGAAUCAGCUAAUUGUUCAGCA---CACAUUUGUGUGGGCAAUUUACGUGAAUG-AAAAUUAUUUUCGUACAGCACGUCUCUUGUCUUCUUCGGCAAUA .....(((((((((...............(((((((((.((---((....))))))))))))).(((((.(((-(((.....))))))....))))).)))))))))........... ( -29.10, z-score = -2.55, R) >droPer1.super_66 57180 102 + 382987 ---------GGUGGUUAAUCGAAUUGUAUGUAUGCUUUAUAGUUUGGAAUCUUGUGUUUGGCACAUGCGACGGGAAAA-------AGCACAGAGCUUGUGCCGUCGUGCUUGGUAAAA ---------...((((..((((((((((.........)))))))))))))).....((((.((..((((((((.(..(-------(((.....)))).).))))))))..)).)))). ( -25.90, z-score = -0.32, R) >consensus AGUGGCGGGCGAGAACAAUUGAAUCAGCUAAUUGUUCCACA___CACAUUUGUGUGGGUAAUUUACGUGAAUGAAAAAUUAUUUUCGUACAGCACGUCUCUUGUCUUCUUCGGGAAUA .......((((.(((((((((.......))))))))).........((((..(((((.....)))))..)))).....................))))(((((.......)))))... (-14.34 = -14.40 + 0.06)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:47:59 2011