| Sequence ID | dm3.chrX |
|---|---|
| Location | 15,458,310 – 15,458,408 |
| Length | 98 |
| Max. P | 0.920128 |

| Location | 15,458,310 – 15,458,408 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 77.25 |
| Shannon entropy | 0.44515 |
| G+C content | 0.41534 |
| Mean single sequence MFE | -24.92 |
| Consensus MFE | -15.37 |
| Energy contribution | -16.90 |
| Covariance contribution | 1.53 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.920128 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
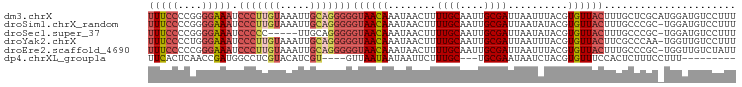
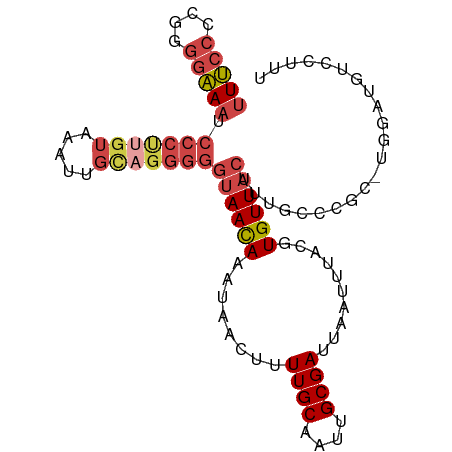
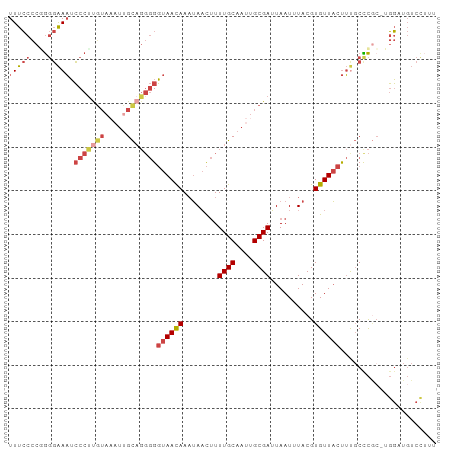
>dm3.chrX 15458310 98 - 22422827 UUUCCCCGGGGAAAUCCCUUGUAAAUUGCAGGGGGUAACAAAUAACUUUUGCAAUUGCGAUUAAUUUACGUGUUACUUUGCUCGCAUGGAUGUCCUUU ..(((.((((.(((.(((((((.....)))))))((((((..(((...((((....)))).....)))..))))))))).))))...)))........ ( -27.00, z-score = -1.57, R) >droSim1.chrX_random 231425 97 - 5698898 UUUCCCCGGGGAAAUCCCUUGUAAAUUGCAGGGGGUAACAAAUAACUUUUGCAAUUGCGAUUAAUAUACGUGUUACUUUGCCCGC-UGGAUGUCCUUU ..(((.((((.(((.(((((((.....)))))))((((((.(((....((((....))))....)))...))))))))).)))).-.)))........ ( -30.00, z-score = -2.29, R) >droSec1.super_37 37581 92 - 454039 UUUCCCCGGGGAAAUCCCCC-----UUGCAGGGGGUAACAAAUAACUUUUGCAAUUGCGAUUAAUAUACGUGUUACUUUGCCCGC-UGGAUGUCCUUU ..(((.((((.(((..((((-----.....))))((((((.(((....((((....))))....)))...))))))))).)))).-.)))........ ( -27.40, z-score = -1.44, R) >droYak2.chrX 9639910 97 - 21770863 UUUCCCCUGGGAAAUCCCUUGUAAAUUGCAGGGGGUAACAAAUAACUUUUGCAAUUGCGAUUAAUUUACGUGUUACUUCGCCCAA-UGGUUGUCCUUU ...((..((((....(((((((.....)))))))((((((..(((...((((....)))).....)))..))))))....)))).-.))......... ( -26.80, z-score = -1.95, R) >droEre2.scaffold_4690 7169317 97 + 18748788 UUUCCCCCGGGAAAUCCCUUGUAAAUUGCAGGGGGUAACAAAUAACUUUUGCAAUUGCGAUUAAUUUACGUGUUACUUUGCCCGC-UGGUUGUCUAUU ...((..((((....(((((((.....)))))))((((((..(((...((((....)))).....)))..))))))....)))).-.))......... ( -27.80, z-score = -2.01, R) >dp4.chrXL_group1a 1043367 82 - 9151740 UUCACUCAACCGAUGGCCUCGUACAUCGU----GUUAAUAAUAAUUCUUUGC---UGCGAAUAAUCUACGUGUUUCCACUCUUUCCUUU--------- ..(((..((((((((........))))).----))).......((((.....---...)))).......))).................--------- ( -10.50, z-score = -1.05, R) >consensus UUUCCCCGGGGAAAUCCCUUGUAAAUUGCAGGGGGUAACAAAUAACUUUUGCAAUUGCGAUUAAUUUACGUGUUACUUUGCCCGC_UGGAUGUCCUUU (((((....))))).(((((((.....)))))))((((((........((((....))))..........))))))...................... (-15.37 = -16.90 + 1.53)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:46:46 2011