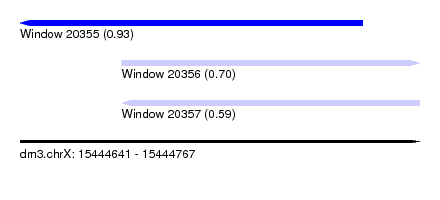

| Sequence ID | dm3.chrX |
|---|---|
| Location | 15,444,641 – 15,444,767 |
| Length | 126 |
| Max. P | 0.925198 |

| Location | 15,444,641 – 15,444,749 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 96.30 |
| Shannon entropy | 0.06270 |
| G+C content | 0.28519 |
| Mean single sequence MFE | -23.68 |
| Consensus MFE | -21.64 |
| Energy contribution | -21.76 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.91 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.31 |
| SVM RNA-class probability | 0.925198 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
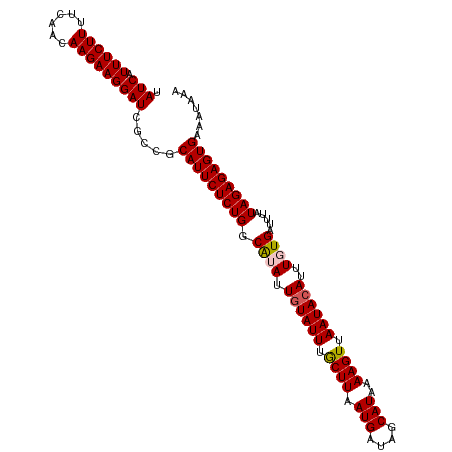
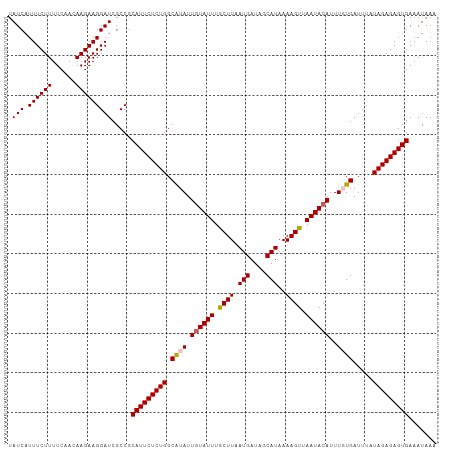
>dm3.chrX 15444641 108 - 22422827 UAUCAUUUCUUUUCAACAAGAAGGAUCGCCACAUUCUCUGGCAUAUUGUAUUUGCUUAAUGAUAUCAUAAAAGUUAAUACAUUUGUGAUUUAUAGAGAGUGAAAUAAA .(((.((((((......))))))))).....(((((((((.((((.((((((.((((.(((....)))..)))).))))))..)))).....)))))))))....... ( -25.40, z-score = -2.86, R) >droSim1.chrX_random 4033937 108 - 5698898 UAUCAUUUCUUUUCAACAAGAAGGAUCGCCGCAUUCUCUGGCAUAUUGUAUUUGCUUAAUGAUAGCAUAAAAGUUAAUACAUUUGUGAUUUAUAGAGAGUGAAAUAAA .(((.((((((......))))))))).....(((((((((.((((.((((((.((((.(((....)))..)))).))))))..)))).....)))))))))....... ( -24.30, z-score = -1.94, R) >droSec1.super_37 19872 108 - 454039 UAUCAUUUCUUUUCAACAAGAAGGAUCGCCGCAUUCUCUGGCAUAUUGUAUUUGCUUAAUGAUAGCAUAAAAGUUAAUACAUUUGUGAUUUAUAGAGAGUGAAAUAAA .(((.((((((......))))))))).....(((((((((.((((.((((((.((((.(((....)))..)))).))))))..)))).....)))))))))....... ( -24.30, z-score = -1.94, R) >droYak2.chrX 9628836 108 - 21770863 UAUCAUUUCUUUUCAAUAAGAAGGAUCGCCGCAUUCUCUGGCGUAUUGUAUUUACUUUAUGAUAGCAUAAAAGUUAAUACAUUUUUGAUUUAUAGAGAGUGAAAUAAA .(((.((((((......))))))))).....(((((((((.((...((((((.((((((((....))).))))).))))))....)).....)))))))))....... ( -23.80, z-score = -2.33, R) >droEre2.scaffold_4690 7157844 108 + 18748788 UAUCAUUUCUUUUCAACAAGAAGGAUCGCCGCAUUCUCUGGCGUAUUUUAUUUACUUUAUGAUAGCAUAAAAGUUAAUACAUUUUUGAUUUAUAGAGAGUGAAAUAAA .(((.((((((......)))))))))(((((.......)))))(((((((((..((((((((...((.(((.((....)).))).))..))))))))))))))))).. ( -20.60, z-score = -1.63, R) >consensus UAUCAUUUCUUUUCAACAAGAAGGAUCGCCGCAUUCUCUGGCAUAUUGUAUUUGCUUAAUGAUAGCAUAAAAGUUAAUACAUUUGUGAUUUAUAGAGAGUGAAAUAAA .(((.((((((......))))))))).....(((((((((.((((.((((((.((((.(((....)))..)))).))))))..)))).....)))))))))....... (-21.64 = -21.76 + 0.12)

| Location | 15,444,673 – 15,444,767 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 86.77 |
| Shannon entropy | 0.20774 |
| G+C content | 0.37029 |
| Mean single sequence MFE | -22.02 |
| Consensus MFE | -18.80 |
| Energy contribution | -18.84 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.85 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.701865 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
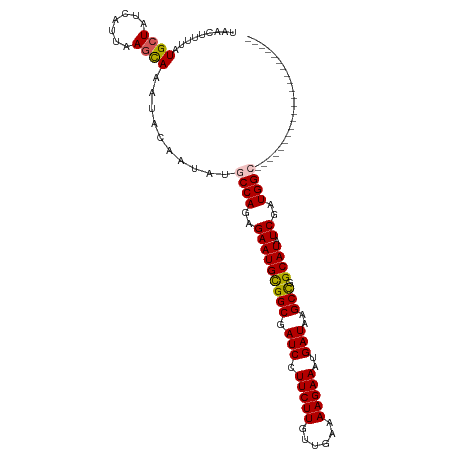
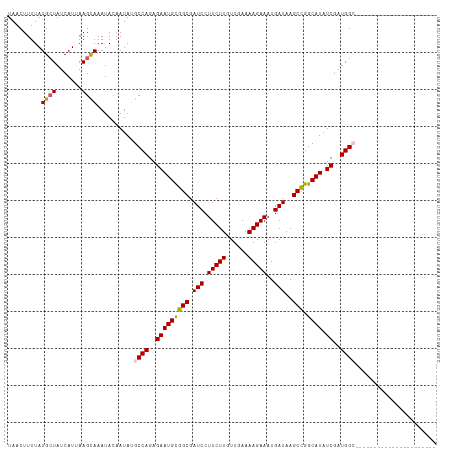
>dm3.chrX 15444673 94 + 22422827 UAACUUUUAUGAUAUCAUUAAGCAAAUACAAUAUGCCAGAGAAUGUGGCGAUCCUUCUUGUUGAAAAGAAAUGAUAAGCCGGCAUAUCGAUGGC---------------------- ...(((..(((....))).)))............((((..(((((((((.(((.(((((......)))))..)))..))).)))).))..))))---------------------- ( -19.70, z-score = -1.06, R) >droSim1.chrX_random 4033969 94 + 5698898 UAACUUUUAUGCUAUCAUUAAGCAAAUACAAUAUGCCAGAGAAUGCGGCGAUCCUUCUUGUUGAAAAGAAAUGAUAAGCCGGCAUAUCGAUGGC---------------------- .........((((.......))))..........((((..(((((((((.(((.(((((......)))))..)))..))).)))).))..))))---------------------- ( -23.50, z-score = -1.92, R) >droSec1.super_37 19904 94 + 454039 UAACUUUUAUGCUAUCAUUAAGCAAAUACAAUAUGCCAGAGAAUGCGGCGAUCCUUCUUGUUGAAAAGAAAUGAUAAGCCGGCAUAUCGAUGGC---------------------- .........((((.......))))..........((((..(((((((((.(((.(((((......)))))..)))..))).)))).))..))))---------------------- ( -23.50, z-score = -1.92, R) >droYak2.chrX 9628868 116 + 21770863 UAACUUUUAUGCUAUCAUAAAGUAAAUACAAUACGCCAGAGAAUGCGGCGAUCCUUCUUAUUGAAAAGAAAUGAUAAGCUGGCAUAUCAAUGGAGAGAAAGAGUGAGAGAUGGAGA ...(((((((.((.((.....(((.......))).(((..(((((((((.(((.(((((......)))))..)))..))).)))).))..)))...)).)).)))))))....... ( -22.60, z-score = -0.89, R) >droEre2.scaffold_4690 7157876 94 - 18748788 UAACUUUUAUGCUAUCAUAAAGUAAAUAAAAUACGCCAGAGAAUGCGGCGAUCCUUCUUGUUGAAAAGAAAUGAUAAGCCGGCAUAUCGAUGGA---------------------- ..(((((.(((....))))))))............(((..(((((((((.(((.(((((......)))))..)))..))).)))).))..))).---------------------- ( -20.80, z-score = -1.61, R) >consensus UAACUUUUAUGCUAUCAUUAAGCAAAUACAAUAUGCCAGAGAAUGCGGCGAUCCUUCUUGUUGAAAAGAAAUGAUAAGCCGGCAUAUCGAUGGC______________________ .........((((.......))))..........((((..(((((((((.(((.(((((......)))))..)))..))).)))).))..))))...................... (-18.80 = -18.84 + 0.04)

| Location | 15,444,673 – 15,444,767 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 86.77 |
| Shannon entropy | 0.20774 |
| G+C content | 0.37029 |
| Mean single sequence MFE | -20.76 |
| Consensus MFE | -17.00 |
| Energy contribution | -16.68 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.587785 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 15444673 94 - 22422827 ----------------------GCCAUCGAUAUGCCGGCUUAUCAUUUCUUUUCAACAAGAAGGAUCGCCACAUUCUCUGGCAUAUUGUAUUUGCUUAAUGAUAUCAUAAAAGUUA ----------------------((...((((((((((((..(((.((((((......))))))))).))).........))))))))).....))...(((....)))........ ( -19.50, z-score = -1.45, R) >droSim1.chrX_random 4033969 94 - 5698898 ----------------------GCCAUCGAUAUGCCGGCUUAUCAUUUCUUUUCAACAAGAAGGAUCGCCGCAUUCUCUGGCAUAUUGUAUUUGCUUAAUGAUAGCAUAAAAGUUA ----------------------((((..((.((((.(((..(((.((((((......))))))))).)))))))))..))))..........((((.......))))......... ( -23.30, z-score = -1.80, R) >droSec1.super_37 19904 94 - 454039 ----------------------GCCAUCGAUAUGCCGGCUUAUCAUUUCUUUUCAACAAGAAGGAUCGCCGCAUUCUCUGGCAUAUUGUAUUUGCUUAAUGAUAGCAUAAAAGUUA ----------------------((((..((.((((.(((..(((.((((((......))))))))).)))))))))..))))..........((((.......))))......... ( -23.30, z-score = -1.80, R) >droYak2.chrX 9628868 116 - 21770863 UCUCCAUCUCUCACUCUUUCUCUCCAUUGAUAUGCCAGCUUAUCAUUUCUUUUCAAUAAGAAGGAUCGCCGCAUUCUCUGGCGUAUUGUAUUUACUUUAUGAUAGCAUAAAAGUUA ............................((((((((((........(((((......)))))((....)).......))))))))))........((((((....))))))..... ( -19.20, z-score = -1.11, R) >droEre2.scaffold_4690 7157876 94 + 18748788 ----------------------UCCAUCGAUAUGCCGGCUUAUCAUUUCUUUUCAACAAGAAGGAUCGCCGCAUUCUCUGGCGUAUUUUAUUUACUUUAUGAUAGCAUAAAAGUUA ----------------------.(((..((.((((.(((..(((.((((((......))))))))).)))))))))..))).(((.......)))((((((....))))))..... ( -18.50, z-score = -1.25, R) >consensus ______________________GCCAUCGAUAUGCCGGCUUAUCAUUUCUUUUCAACAAGAAGGAUCGCCGCAUUCUCUGGCAUAUUGUAUUUGCUUAAUGAUAGCAUAAAAGUUA ...........................(((((((((((........(((((......)))))((....)).......)))))))))))............................ (-17.00 = -16.68 + -0.32)

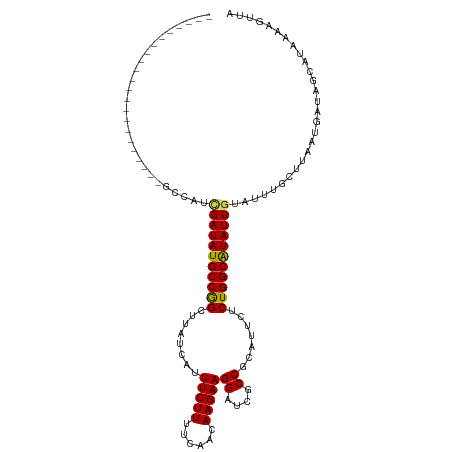
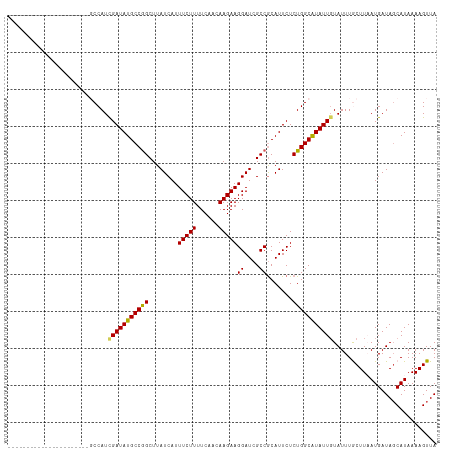
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:46:45 2011