
| Sequence ID | dm3.chrX |
|---|---|
| Location | 15,261,079 – 15,261,198 |
| Length | 119 |
| Max. P | 0.754950 |

| Location | 15,261,079 – 15,261,179 |
|---|---|
| Length | 100 |
| Sequences | 11 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 65.94 |
| Shannon entropy | 0.70179 |
| G+C content | 0.33885 |
| Mean single sequence MFE | -19.46 |
| Consensus MFE | -6.13 |
| Energy contribution | -5.95 |
| Covariance contribution | -0.18 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.31 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.754950 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 15261079 100 - 22422827 UUAUCGAUUACCUUGUCAUUUAUCACCGAUUCCGUGUGAUUGACUUAAUUAAAUGUGUAACCAUAUUUAAUGUUGACCA----ACCUGAUA---GUCCGCAAAAAAU-- ((((((........((((...(((((((....)).)))))))))...((((((((((....))))))))))........----...)))))---)............-- ( -18.60, z-score = -2.44, R) >droSim1.chrX 11780041 100 - 17042790 UUAUCGAUUACCUUGUCAUUUAUCACCGAUUCCGUGUGAUUGACUUAAUUAAAUGUGUAACCAUAUUUAAUGUUGACCA----ACUGGAUG---GGCCGCAAAAAGU-- ...(((((......((((...(((((((....)).)))))))))...((((((((((....)))))))))))))))(((----......))---)............-- ( -20.00, z-score = -0.78, R) >droSec1.super_56 146070 99 - 192059 UUAUCGAUUACCUUGUCAUUUAUCACCGAUUCCGUGUGAUUGACUUAAUUAAAUGUGU-ACCAUAUUUAAUGUUGACCA----ACUGGAUA---GGCCGCAAAAAGU-- ...(((((......((((...(((((((....)).)))))))))...((((((((((.-..)))))))))))))))((.----...))...---.............-- ( -17.90, z-score = -0.65, R) >droYak2.chrX 9457845 105 - 21770863 UUAUCGAUUACCUUGUCAUUUAUCACCGACUCCGUUUGAUUGACUUAAUUAAAUGUGUAACCAUAUUUAAUGUCGACCA----ACAGGAUGGUUGGCCACAAAAAAAUA .....(((......)))........((((((((..(((.(((((...((((((((((....))))))))))))))).))----)..)))..)))))............. ( -22.60, z-score = -1.79, R) >droEre2.scaffold_4690 6986640 100 + 18748788 UUAUCGAUUACCUUUUCAUUUAUCACCGAUUCCGUUUGAUUGACUUAAUUAAAUGUGUAACCAUAUUUAAUGUUGACCA----ACAGGAUA---GGCGGCAAAAAAA-- ..................(((..(.((...(((..(((.(..((...((((((((((....))))))))))))..).))----)..)))..---)).)..)))....-- ( -18.90, z-score = -1.39, R) >droAna3.scaffold_13117 1478137 84 - 5790199 UUAUCGAUUACGCUGUCAUUUAUCGCCG-----GAUUGAUUGACAUAAUUAAAUGUGUUACCAUAUUUAAUGCUUCCCC---GACAGUAAAA----------------- ...........(((((((((.(((....-----))).))).......((((((((((....))))))))))........---))))))....----------------- ( -15.90, z-score = -1.66, R) >dp4.chrXL_group1e 12198655 99 - 12523060 ---UUAGGGACACAGGUCCAUUUUGCC-----CGACUGAUUGACAUAAUUAAAUGUGUGACCAUAUUUAAUGUUGGCCUAACAAUAACAGAACAAAGCAUAAAAAGC-- ---....((((....))))....(((.-----...(((..........(((((((((....)))))))))(((((......))))).)))......)))........-- ( -17.30, z-score = -0.28, R) >droPer1.super_14 759904 99 + 2168203 ---UUAUGGACACAGGUCCAUAUCGCC-----CGACUGAUUGACAUAAUUAAAUGUGUGACCAUAUUUAAUGUUGGCCUAACAAUAACAGAACAAAGCAUAAAAAGC-- ---.(((((((....)))))))..((.-----...(((..........(((((((((....)))))))))(((((......))))).)))......)).........-- ( -20.00, z-score = -1.58, R) >droVir3.scaffold_12928 7283970 97 - 7717345 UUAUCGAUUGUUAUGGUAUUUAUCAGC------ACUAGAUUGACAUGAUUAAAUAUGCAGUUGUAUUAAAUAUUGCACC----AUAACAAAA-CGAUAGCCAAACACC- ((((((.((((((((((((((((((.(------(......))...))).))))))((((((..........))))))))----)))))))..-)))))).........- ( -22.50, z-score = -3.19, R) >droMoj3.scaffold_6473 14157583 97 + 16943266 UUAUCGAUUGUUAUGGUAUUUAUCACC------ACCGGAUUGACAUUAUUAAAUAUGUAACCAUAUUUAAUAUUGCACC----AUAACAAAA-CGGUAGCCAAACAAA- ((((((.((((((((((........((------...))...(.((.(((((((((((....))))))))))).))))))----)))))))..-)))))).........- ( -24.80, z-score = -4.09, R) >droGri2.scaffold_15081 847849 97 + 4274704 UUAUCGAUUGUUACGGUAUUUUUCACC------ACUGGAUUGACAUGAUUAAAUAUGUAACCAUAUUAAAUACUGUACC----AUGACAAAA-CGACUGCCACACAUC- ...(((.((((((.(((((........------..(((....(((((......)))))..)))...........)))))----.))))))..-)))............- ( -15.60, z-score = -0.69, R) >consensus UUAUCGAUUACCUUGUCAUUUAUCACCG____CGUUUGAUUGACAUAAUUAAAUGUGUAACCAUAUUUAAUGUUGACCA____ACAACAUAA__GGCCGCAAAAAAU__ ...............................................((((((((((....))))))))))...................................... ( -6.13 = -5.95 + -0.18)


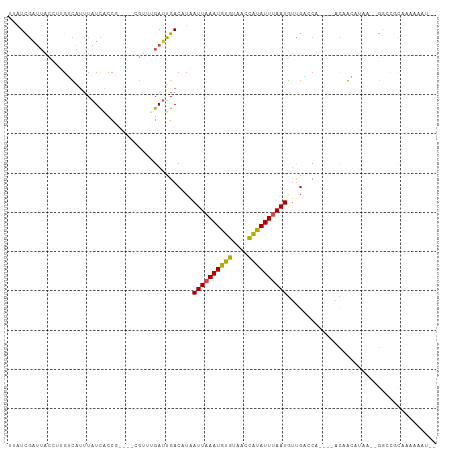
| Location | 15,261,107 – 15,261,198 |
|---|---|
| Length | 91 |
| Sequences | 12 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 62.69 |
| Shannon entropy | 0.73618 |
| G+C content | 0.35692 |
| Mean single sequence MFE | -16.82 |
| Consensus MFE | -6.26 |
| Energy contribution | -6.02 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.40 |
| Mean z-score | -0.94 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.521616 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 15261107 91 - 22422827 ----------------CAUCAGCAGCAUUGUCCGC-UUAUCGAUUACCUUGUCAUUUAUCACCGAUUCCGUGUGAUUGACUUAAUUAAAUGUGUAACCAUAUUUAAUG ----------------.(((...(((.......))-)....)))......((((...(((((((....)).)))))))))...((((((((((....)))))))))). ( -17.00, z-score = -1.81, R) >droSim1.chrX 11780069 91 - 17042790 ----------------CAUCAGCAGCAUUGUCCGC-UUAUCGAUUACCUUGUCAUUUAUCACCGAUUCCGUGUGAUUGACUUAAUUAAAUGUGUAACCAUAUUUAAUG ----------------.(((...(((.......))-)....)))......((((...(((((((....)).)))))))))...((((((((((....)))))))))). ( -17.00, z-score = -1.81, R) >droSec1.super_56 146098 90 - 192059 ----------------CAUCAGCAGCAUUGUCCGC-UUAUCGAUUACCUUGUCAUUUAUCACCGAUUCCGUGUGAUUGACUUAAUUAAAUGUGU-ACCAUAUUUAAUG ----------------.(((...(((.......))-)....)))......((((...(((((((....)).)))))))))...((((((((((.-..)))))))))). ( -15.90, z-score = -1.37, R) >droYak2.chrX 9457878 91 - 21770863 ----------------CAUCAACAGCAUUGUCUGC-UUAUCGAUUACCUUGUCAUUUAUCACCGACUCCGUUUGAUUGACUUAAUUAAAUGUGUAACCAUAUUUAAUG ----------------.......((((.....)))-)..(((((((....(((..........)))......)))))))....((((((((((....)))))))))). ( -13.70, z-score = -1.33, R) >droEre2.scaffold_4690 6986668 91 + 18748788 ----------------CAUCAGCAGCAUUGUCUGG-UUAUCGAUUACCUUUUCAUUUAUCACCGAUUCCGUUUGAUUGACUUAAUUAAAUGUGUAACCAUAUUUAAUG ----------------.........(((((..(((-((((.((........))...............((((((((((...)))))))))).)))))))....))))) ( -12.50, z-score = -0.31, R) >droAna3.scaffold_13117 1478154 103 - 5790199 CGAAUCCCAUUUGCAGGCGGCGGGGCAAAUGGCCAGUUAUCGAUUACGCUGUCAUUUAUCGCCGGAU-----UGAUUGACAUAAUUAAAUGUGUUACCAUAUUUAAUG ..(((((.....(....)(((((...(((((((.(((..........)))))))))).)))))))))-----)..........((((((((((....)))))))))). ( -25.00, z-score = -0.05, R) >dp4.chrXL_group1e 12198690 83 - 12523060 -------------------CAUCUACUCAAUUGCC-UGGUUAGGGACACAGGUCCAUUUUGCCCGAC-----UGAUUGACAUAAUUAAAUGUGUGACCAUAUUUAAUG -------------------.......(((((((.(-.(((...((((....)))).....))).).)-----.))))))....((((((((((....)))))))))). ( -18.60, z-score = -1.05, R) >droPer1.super_14 759939 83 + 2168203 -------------------CAUCUACUCAAUUGCC-UGGUUAUGGACACAGGUCCAUAUCGCCCGAC-----UGAUUGACAUAAUUAAAUGUGUGACCAUAUUUAAUG -------------------.......(((((((..-(((.(((((((....)))))))....)))..-----)))))))....((((((((((....)))))))))). ( -21.50, z-score = -2.39, R) >droWil1.scaffold_180702 2773992 97 + 4511350 -----CACCUGGUGUGGUUCCUGGGCCUAGUCGGUGUUAACAAUUACUGUAUUAUUUAUUGCCCAA------UGAUUGACAUGAUUAAAUGUGUAACCAUAUUUGAUG -----...(..((((((((.((((((.(((((((((........))))).))))......))))).------......((((......))))).))))))))..)... ( -22.20, z-score = -0.50, R) >droVir3.scaffold_12928 7284001 70 - 7717345 --------------------------------CCAGUUAUCGAUUGUUAUGGUAUUUAUCAGCACU------AGAUUGACAUGAUUAAAUAUGCAGUUGUAUUAAAUA --------------------------------........(((((((....((((((((((.((..------....))...))).))))))))))))))......... ( -9.80, z-score = 0.37, R) >droMoj3.scaffold_6473 14157614 86 + 16943266 --------------UAAUCGAUAGGAC--GUCCUAGUUAUCGAUUGUUAUGGUAUUUAUCACCACC------GGAUUGACAUUAUUAAAUAUGUAACCAUAUUUAAUA --------------(((((((((((..--..))))....)))))))...((((.......))))..------..........(((((((((((....))))))))))) ( -18.30, z-score = -1.42, R) >droGri2.scaffold_15081 847880 76 + 4274704 ------------------------GCC--GUCCUAGUUAUCGAUUGUUACGGUAUUUUUCACCACU------GGAUUGACAUGAUUAAAUAUGUAACCAUAUUAAAUA ------------------------(((--((..(((((...)))))..)))))............(------((....(((((......)))))..)))......... ( -10.30, z-score = 0.33, R) >consensus ________________CAUCAGCAGCAUUGUCCGC_UUAUCGAUUACCUUGUCAUUUAUCACCGAUU_____UGAUUGACAUAAUUAAAUGUGUAACCAUAUUUAAUG ...................................................................................((((((((((....)))))))))). ( -6.26 = -6.02 + -0.25)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:46:02 2011