| Sequence ID | dm3.chrX |
|---|---|
| Location | 15,217,695 – 15,217,793 |
| Length | 98 |
| Max. P | 0.769206 |

| Location | 15,217,695 – 15,217,793 |
|---|---|
| Length | 98 |
| Sequences | 7 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 65.69 |
| Shannon entropy | 0.61528 |
| G+C content | 0.56377 |
| Mean single sequence MFE | -36.71 |
| Consensus MFE | -16.32 |
| Energy contribution | -16.70 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.72 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.769206 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
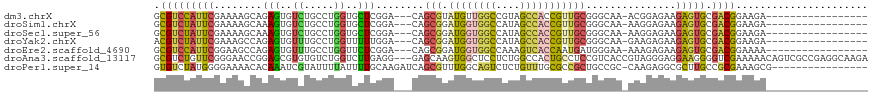
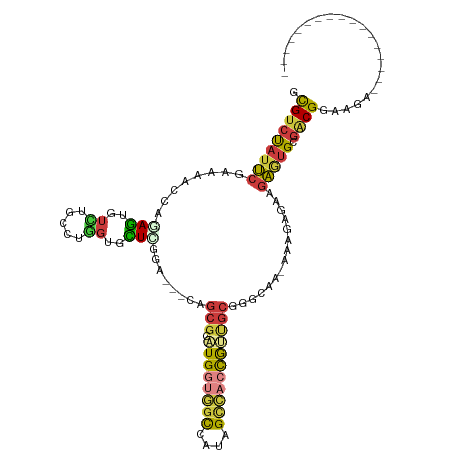
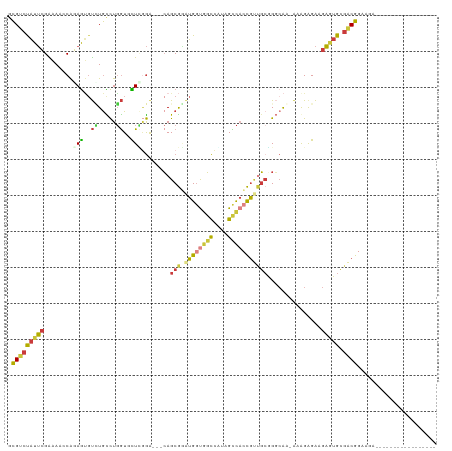
>dm3.chrX 15217695 98 + 22422827 GCGUCCAUUCGAAAAGCAGAGUGUCUGCCUGGUGCUCGGA---CAGCGUAUGUUGGCCGUAGCCACCGUUGCGGGCAA-ACGGAGAAGAGUGCGACGGAAGA----------------- .(((((((((......(....(((((((.((((((((((.---((((....)))).))).)).)))))..))))))).-...)....))))).)))).....----------------- ( -37.70, z-score = -1.41, R) >droSim1.chrX 11740706 98 + 17042790 GCGUCUAUUCGAAAAGCAAAGUGUCUGCCUGGUGCUCGGA---CAGCGGAUGGUGGCCAUAGCCACCGUUGCGGGCAA-AAGGAGAAGAGUGCGACGGAAGA----------------- ...(((.((((....(((.....(((.(((..((((((..---((((((.((((.......)))))))))))))))).-.))))))....)))..)))))))----------------- ( -36.80, z-score = -1.81, R) >droSec1.super_56 105453 98 + 192059 GCGUCUAUUCGAAAAGCAAAGUGUCUGCCUGGUGCUCGGA---CAGCGGAUGGUGGCCAUAGCCACCGUUGCGGGCAA-AAGGAGAAGAGUGCGACGGAAGA----------------- ...(((.((((....(((.....(((.(((..((((((..---((((((.((((.......)))))))))))))))).-.))))))....)))..)))))))----------------- ( -36.80, z-score = -1.81, R) >droYak2.chrX 9417713 98 + 21770863 ACGUCUAUUCGAAAGCCAGAGUGUUUGCCUGGUUUUUGGA---CAGCGGAUGGUGGCCAUAGCCACCGUUGCGGGCAA-GAAGAGAAGAGUGCGACGGAAGA----------------- .((((..((((((((((((.((....))))))))))))))---..(((.((((((((....)))))))))))..(((.-...........))))))).....----------------- ( -38.10, z-score = -3.06, R) >droEre2.scaffold_4690 6945074 98 - 18748788 GCGUCCAUUCGGAAGCCAGAGUGUUUGCCUGGUUCUCGGA---CAGCGGAUGGUGGCCAAAGUCACCAAUGAUGGGAA-AAAGAGAAGAGUGCGACGGAAAA----------------- .((((((((((..((((((.((....))))))))..)...---((.((..(((((((....))))))).)).))....-........))))).)))).....----------------- ( -33.40, z-score = -1.83, R) >droAna3.scaffold_13117 1430622 116 + 5790199 GCGUCUGUUCGGGAACCGGAGCGUGUGUCUGGUCUUGAGG---GAGCAAGUGGCUCCUCUGGCCACUGCCUCCGUCACCGUAGGGAGGAAGGGGUCGAAAAACAGUCGCCGAGGCAAGA (((.((((((((((.(((((.......)))))))))))((---((((.....)))))).(((((.((.(((((.(......).))))).)).)))))....)))).))).......... ( -45.20, z-score = -0.04, R) >droPer1.super_14 715452 102 - 2168203 GUGUCUAUGGGGAAAACACAAAUCGUAUUUUAUUUUGCAAGAUCAGCGUUUGGCAGUCUCUGUUUGCGCCGCUGCCGC-CAAGAGGCGCUUGCCGCGAAAGCG---------------- ((((((...((..((((.(..((((((........)))..)))..).))))((((((...((....))..)))))).)-)...))))))....(((....)))---------------- ( -29.00, z-score = 0.82, R) >consensus GCGUCUAUUCGAAAACCAGAGUGUCUGCCUGGUGCUCGGA___CAGCGGAUGGUGGCCAUAGCCACCGUUGCGGGCAA_AAAGAGAAGAGUGCGACGGAAGA_________________ .(((((((((........(((..((.....))..)))........(((.((((((((....)))))))))))...............))))).))))...................... (-16.32 = -16.70 + 0.38)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:45:57 2011