| Sequence ID | dm3.chrX |
|---|---|
| Location | 15,192,623 – 15,192,746 |
| Length | 123 |
| Max. P | 0.904545 |

| Location | 15,192,623 – 15,192,746 |
|---|---|
| Length | 123 |
| Sequences | 6 |
| Columns | 129 |
| Reading direction | reverse |
| Mean pairwise identity | 76.83 |
| Shannon entropy | 0.42850 |
| G+C content | 0.56017 |
| Mean single sequence MFE | -42.40 |
| Consensus MFE | -28.35 |
| Energy contribution | -29.83 |
| Covariance contribution | 1.48 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.18 |
| SVM RNA-class probability | 0.904545 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
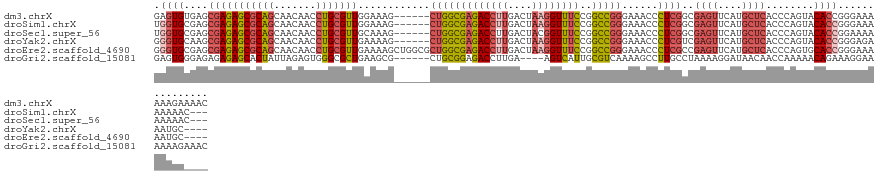
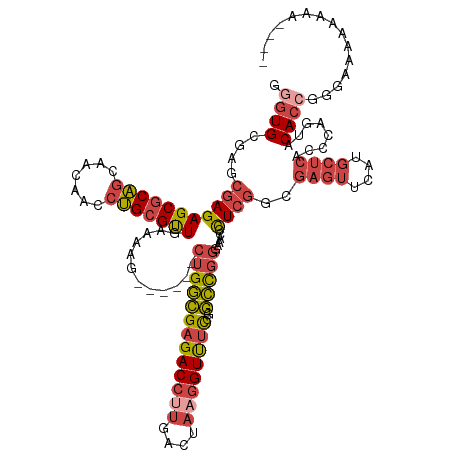
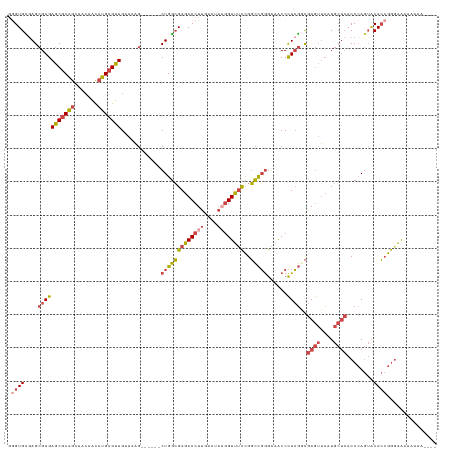
>dm3.chrX 15192623 123 - 22422827 GAGUGUGAGCGAGAGCGCAGCAACAACCUGCGUUGGAAAG------CUGGCGAGACCUUGACUAAGGUUUCCGGCCGGGAAACCCUCGGCGAGUUCAUGCUCACCCAGUACACCGGGAAAAAAGAAAAC ((((((((((....((.((((..((((....))))....)------)))))((((((((....))))))))..((((((.....))))))..)))))))))).(((........)))............ ( -48.20, z-score = -3.13, R) >droSim1.chrX 11715291 120 - 17042790 UGGUGCGAGCGAGAGCGCAGCAACAACCUGCGUUGGAAAG------CUGGCGAGACCUUGACUAAGGUUUCCGGCCGGGAAACCCUCGGCGAGUUCAUGCUCACCCAGUACACCGGGAAAAAAAAC--- ....((.(((...(((((((.......))))))).....)------)).))((((((((....))))))))..((((((.....))))))((((....)))).(((........))).........--- ( -43.70, z-score = -1.58, R) >droSec1.super_56 80847 120 - 192059 UGGUGCGAGCGAGAGCGCAGCAACAACCUGCGUUGCAAAG------CUGGCGAGACCUUGACUACGGUUUCCGGCCGGGAAACCCUCGGCGAGUUCAUGCUCACCCAGUACACCGGAAAAAAAAAC--- .((((.(.((....)).).(((((.......)))))...(------((((.((((((........))))))..((((((.....))))))((((....))))..))))).))))............--- ( -42.80, z-score = -1.59, R) >droYak2.chrX 9393815 119 - 21770863 GGGUGCAAGCGAGAGCGCAGCAACAACCUGCGUUGAAAAG------CUGGCGAGACCUUGACUAAGGUUUCCGGCCGGGAAACCCUCGUCGAGUUCAUGCUCACCCAGUACACCGGGAGAAAUGC---- ((((((..((((((((((((.......)))))))......------..(((((((((((....))))))))..)))((....))))))).((((....)))).....)))).))...........---- ( -43.10, z-score = -1.39, R) >droEre2.scaffold_4690 6921041 125 + 18748788 GGGUGCGAGCGAGAGCGCAGCAACAACCUGCGUUGAAAAGCUGGCGCUGGCGAGACCUUGACUAAGGUUUCCGGCCGGGAAACCCUCGCCGAGUUCAUGCUCACCCAGUGCACCGGGAAAAAUGC---- .((((((.((((((((((.(((......)))(((....)))..)))))(((((((((((....))))))))..)))((....))))))))((((....)))).......)))))...........---- ( -52.80, z-score = -2.18, R) >droGri2.scaffold_15081 1542623 119 + 4274704 GAGUGGGAGAGAGAGCACUAUUAGAGUGGGCGCUGAAGCG------CUGCGGAGACCUUGA----AGUCAUUGCGUCAAAAGCCUUGCCUAAAAGGAUAACAACCAAAAACAGAAAGGAAAAAAGAAAC ...((((...(((.((............(((((....)))------))((((.(((.....----.))).)))).......))))).))))...((.......))........................ ( -23.80, z-score = 0.31, R) >consensus GGGUGCGAGCGAGAGCGCAGCAACAACCUGCGUUGAAAAG______CUGGCGAGACCUUGACUAAGGUUUCCGGCCGGGAAACCCUCGGCGAGUUCAUGCUCACCCAGUACACCGGGAAAAAAAA____ .((((....(((((((((((.......)))))))............(((((((((((((....))))))))..)))))......))))..((((....))))........))))............... (-28.35 = -29.83 + 1.48)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:45:53 2011