
| Sequence ID | dm3.chrX |
|---|---|
| Location | 15,175,084 – 15,175,177 |
| Length | 93 |
| Max. P | 0.587535 |

| Location | 15,175,084 – 15,175,177 |
|---|---|
| Length | 93 |
| Sequences | 10 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 79.35 |
| Shannon entropy | 0.43242 |
| G+C content | 0.34765 |
| Mean single sequence MFE | -12.94 |
| Consensus MFE | -10.27 |
| Energy contribution | -10.18 |
| Covariance contribution | -0.09 |
| Combinations/Pair | 1.10 |
| Mean z-score | -0.70 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.587535 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 15175084 93 - 22422827 --UUGAUUAAUGAAGCAGCUGAGACAACAUCAGCAAUCAAAUUAAUUUCAGUUACAUGAGCGGAUUAUAAAGAAACGACAACAACAGCAACGAAC------- --............((.(((((.......))))).....((((...((((......))))..))))....................)).......------- ( -11.70, z-score = -0.72, R) >droEre2.scaffold_4690 6904087 93 + 18748788 --UUGAUUAAUGAAGCAGCUGAGACAACAUCAGCAAUCAAAUUAAUUUCAGUUACAUGAGCGGAUUAUGAAGAAGCGACAACAACAGCAACGAAC------- --........((..((.(((((.......)))))..(((((((...((((......))))..)))).)))....))..))...............------- ( -14.40, z-score = -0.81, R) >droYak2.chrX 9376055 100 - 21770863 --UUGAUUAAUGAAGCAGCUGAGACAACAUCAGCAAUCAAAUUAAUUUCAGUUACAUGAGCGGAUUAUAAAGAAACGACAACAACAGCAAGCAGCAAGCAAC --............((.(((((.......))))).....((((...((((......))))..))))....................))..((.....))... ( -12.90, z-score = -0.03, R) >droSec1.super_56 63598 93 - 192059 --UUGAUUAAUGAAGCAGCUGAGACAACAUCAGCAAUCAAAUUAAUUUCAGUUACAUGAGCGGAUUAUAAAGAAACGACAACAACAGCAACGAAC------- --............((.(((((.......))))).....((((...((((......))))..))))....................)).......------- ( -11.70, z-score = -0.72, R) >droSim1.chrX_random 3977035 93 - 5698898 --UUGAUUAAUGAAGCAGCUGAGACAACAUCAGCAAUCAAAUUAAUUUCAGUUACAUGAGCGGAUUAUAAAGAAACGACAACAACCGCAACGAAC------- --..(((((((......(((((.......)))))......)))))))(((......)))((((.....................)))).......------- ( -14.80, z-score = -1.84, R) >dp4.chrXL_group1a 7536439 92 + 9151740 --UUGAUUAAUGAAGCAGCUGAGACAACAUCAGCAAUCAAAUUAAUUUCAGUUGCAUGAGCGGAUUAUAGAAU--UUAACAACAAAAGAAGCAAAA------ --.(((((..(.(.((((((((((.....................)))))))))).).)...)))))......--.....................------ ( -12.70, z-score = -0.02, R) >droVir3.scaffold_13042 2132686 79 - 5191987 --UUGAUAAAUGAAGCAGCUGAGACAACAUCAGCAAUCAAAUUAAUUUCAGCUACAUGAGCGGAUUAUGCACGCCGAAAAG--------------------- --............((((((((.......)))))........(((((...(((.....))).))))))))...........--------------------- ( -13.60, z-score = -0.43, R) >droMoj3.scaffold_6359 2035677 88 - 4525533 UUUUGAUUAAUGAAGCAGCUGAGACAACAUCAGCAAUCAAAUUAAUUUUAGUUAUAUGAGCGGAUUAUAUUAAAGCACAGACACAAUA-------------- (((((((((((...((.(((((.......)))))..(((...((((....))))..)))))..)))).))))))).............-------------- ( -12.80, z-score = -0.81, R) >droGri2.scaffold_14853 3426333 100 - 10151454 --UUGAUUAAUGCAGCAGCUGAGACAACAUCAGCAAUCAAAUUAAUUUCAGUUAUAUGAGCGGAUUAAACAACAACAAAUGUAAGCAAAACCAAAAAAAAAC --(((.(((((.(.((.(((((.......))))).............(((......)))))).))))).))).............................. ( -14.20, z-score = -1.16, R) >droAna3.scaffold_13117 1391784 79 - 5790199 --UUGAUUAAUGAAGCAGCUGAGACAACAUCAGCAAUCAAAUUAAUUUCAGUUACAUGAGCGGAUUACAGAAAAAAGACGA--------------------- --..(((((((......(((((.......)))))......)))))))(((......))).((..((........))..)).--------------------- ( -10.60, z-score = -0.51, R) >consensus __UUGAUUAAUGAAGCAGCUGAGACAACAUCAGCAAUCAAAUUAAUUUCAGUUACAUGAGCGGAUUAUAAAGAAACGACAACAACAGCAACGAAC_______ ..............((.(((((.......))))).............(((......)))))......................................... (-10.27 = -10.18 + -0.09)

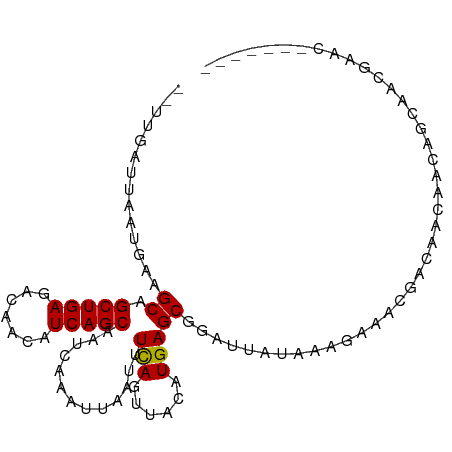

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:45:51 2011