
| Sequence ID | dm3.chrX |
|---|---|
| Location | 15,162,630 – 15,162,733 |
| Length | 103 |
| Max. P | 0.756099 |

| Location | 15,162,630 – 15,162,733 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 77.71 |
| Shannon entropy | 0.35338 |
| G+C content | 0.45148 |
| Mean single sequence MFE | -21.28 |
| Consensus MFE | -12.64 |
| Energy contribution | -16.40 |
| Covariance contribution | 3.76 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.756099 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
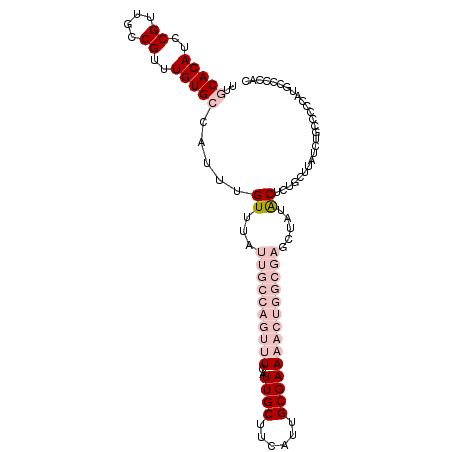
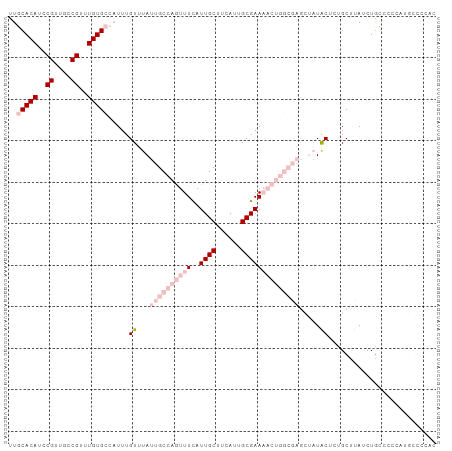
>dm3.chrX 15162630 103 + 22422827 UUGCACAUCCGUUGCCGUUUGUGCCAUUUGUUUAUUGCCAGUUUUAUUGCUUCAUUGCGAAAACUGGCGAGCUAUACUCUGCUUAUCUACCCCCAUGCCCCAC ..(((((..((....))..)))))............((((((((..((((......))))))))))))((((........))))................... ( -27.80, z-score = -3.92, R) >droPer1.super_53 112710 78 + 525408 CUCCACAUUCGUUGUCGUUUGUG---UUUGUUUAUUGCCAAUUUCAUUGCUUCAUUGCGAA-------------UGCUACGCUUAUCUGGACCA--------- .((((.((.....(.(((..(((---(((((.....((..........))......)))))-------------))).))))..)).))))...--------- ( -11.50, z-score = 0.21, R) >dp4.chrXL_group1a 7509940 78 - 9151740 CUCCACAUUCGUUGUCGUUUGUG---UUUGUUUAUUGCCAAUUUCAUUGCUUCAUUGCGAA-------------UGCUACGCUUAUCUGGACCA--------- .((((.((.....(.(((..(((---(((((.....((..........))......)))))-------------))).))))..)).))))...--------- ( -11.50, z-score = 0.21, R) >droSec1.super_56 52200 103 + 192059 UUGCACAUCCGUUGCCGUUUGUGCCAUUUGUUUAUUGCCAGUUUCAUUGCUUCAUUGCGAAAACUGGCGAGCUAUACUCUGCUUAUCUGCCCCCAUGCCCCAC ..(((((..((....))..)))))............((((((((..((((......))))))))))))((((........))))................... ( -27.80, z-score = -3.10, R) >droSim1.chrX 11700202 103 + 17042790 UUGCACAUCCGUUGCCGUUUGUGCCAUUUGUUUAUUGCCAGUUUCAUUGCUUCAUUGCGAAAACUGGCGAGCUAUACUCUGCUUAUCUGCCCCCAUGCCCCAC ..(((((..((....))..)))))............((((((((..((((......))))))))))))((((........))))................... ( -27.80, z-score = -3.10, R) >consensus UUGCACAUCCGUUGCCGUUUGUGCCAUUUGUUUAUUGCCAGUUUCAUUGCUUCAUUGCGAAAACUGGCGAGCUAUACUCUGCUUAUCUGCCCCCAUGCCCCAC ..(((((..((....))..))))).....((...((((((((((..((((......)))))))))))))).....)).......................... (-12.64 = -16.40 + 3.76)


| Location | 15,162,630 – 15,162,733 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 77.71 |
| Shannon entropy | 0.35338 |
| G+C content | 0.45148 |
| Mean single sequence MFE | -24.74 |
| Consensus MFE | -12.76 |
| Energy contribution | -15.80 |
| Covariance contribution | 3.04 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.563471 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
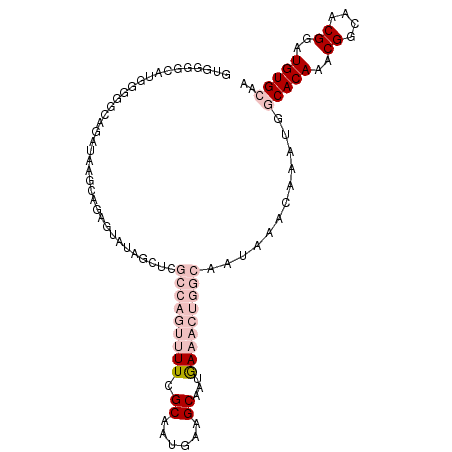
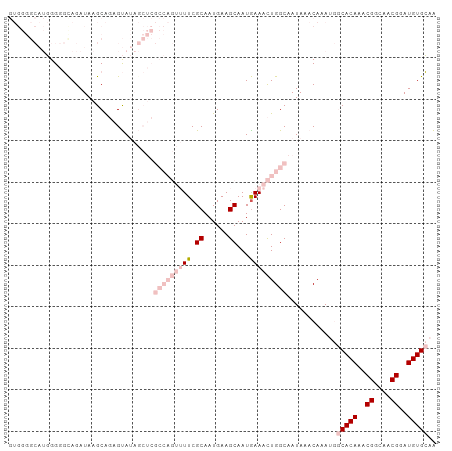
>dm3.chrX 15162630 103 - 22422827 GUGGGGCAUGGGGGUAGAUAAGCAGAGUAUAGCUCGCCAGUUUUCGCAAUGAAGCAAUAAAACUGGCAAUAAACAAAUGGCACAAACGGCAACGGAUGUGCAA ......(((..(..((........(((.....)))(((((((((.((......))...)))))))))..))..)..)))(((((..((....))..))))).. ( -33.10, z-score = -3.70, R) >droPer1.super_53 112710 78 - 525408 ---------UGGUCCAGAUAAGCGUAGCA-------------UUCGCAAUGAAGCAAUGAAAUUGGCAAUAAACAAA---CACAAACGACAACGAAUGUGGAG ---------...((((.((...(((..((-------------((.((......))))))...(((........))).---...........))).)).)))). ( -11.60, z-score = 0.07, R) >dp4.chrXL_group1a 7509940 78 + 9151740 ---------UGGUCCAGAUAAGCGUAGCA-------------UUCGCAAUGAAGCAAUGAAAUUGGCAAUAAACAAA---CACAAACGACAACGAAUGUGGAG ---------...((((.((...(((..((-------------((.((......))))))...(((........))).---...........))).)).)))). ( -11.60, z-score = 0.07, R) >droSec1.super_56 52200 103 - 192059 GUGGGGCAUGGGGGCAGAUAAGCAGAGUAUAGCUCGCCAGUUUUCGCAAUGAAGCAAUGAAACUGGCAAUAAACAAAUGGCACAAACGGCAACGGAUGUGCAA ...(((((((...((......))....))).))))(((((((((.((......))...)))))))))............(((((..((....))..))))).. ( -33.70, z-score = -3.18, R) >droSim1.chrX 11700202 103 - 17042790 GUGGGGCAUGGGGGCAGAUAAGCAGAGUAUAGCUCGCCAGUUUUCGCAAUGAAGCAAUGAAACUGGCAAUAAACAAAUGGCACAAACGGCAACGGAUGUGCAA ...(((((((...((......))....))).))))(((((((((.((......))...)))))))))............(((((..((....))..))))).. ( -33.70, z-score = -3.18, R) >consensus GUGGGGCAUGGGGGCAGAUAAGCAGAGUAUAGCUCGCCAGUUUUCGCAAUGAAGCAAUGAAACUGGCAAUAAACAAAUGGCACAAACGGCAACGGAUGUGCAA ...................................(((((((((.((......))...)))))))))............(((((..((....))..))))).. (-12.76 = -15.80 + 3.04)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:45:47 2011