
| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,920,905 – 14,921,003 |
| Length | 98 |
| Max. P | 0.657008 |

| Location | 14,920,905 – 14,921,003 |
|---|---|
| Length | 98 |
| Sequences | 7 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 63.04 |
| Shannon entropy | 0.65718 |
| G+C content | 0.45822 |
| Mean single sequence MFE | -22.67 |
| Consensus MFE | -7.15 |
| Energy contribution | -7.36 |
| Covariance contribution | 0.21 |
| Combinations/Pair | 1.54 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.32 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.657008 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14920905 98 + 22422827 ----CCCACCC-CCUCGCUCCGCCACUUAUCGCUUCUUUGUGAACGAUUGG---UUUUGAG--------UAUGUAUGCC------GCAUGUGUCGUCAUCUUAUAUGCAUAUACAGGAAU ----.((....-....((((.((((...((((.(((.....))))))))))---)...)))--------).((((((..------(((((((.........))))))).))))))))... ( -24.30, z-score = -1.90, R) >droSim1.chrX 11507598 93 + 17042790 ----CCCACCC-CCUCGCCCCGCCACUUAUCGCUUCUUUGUGAGCGAUUGC---UUUUGAG--------UAUGUGCGCG------GCAUGUGUCGUCAUCUUAUAGAUACAGAAU----- ----.......-..(((((.(((((...(((((((......)))))))(((---(....))--------)))).))).)------)).((((((...........))))))))..----- ( -24.60, z-score = -2.08, R) >droSec1.super_32 375640 93 + 562834 ----CCCACCC-CCUCGCCCCGCCACUUAUCGCUUCUUUGUGAGCGAUUGC---UUUUGAG--------UAUGUGCGCG------GCAUGUGUCGUCAUCUUAUAGAUACAGAAU----- ----.......-..(((((.(((((...(((((((......)))))))(((---(....))--------)))).))).)------)).((((((...........))))))))..----- ( -24.60, z-score = -2.08, R) >droEre2.scaffold_4690 6665096 101 - 18748788 ----CCCACCC-CCUCCUUCUGCCACUUAUCGCUUCUUUGUGAGCGAUUGA---AUUCGAGAAUUCGAGUAUGUGCGCC------GCAUGUGUCAUCAUCUUAUAGAUACAGAAU----- ----.......-.....(((((((((..(((((((......)))))))...---(((((......)))))..))).)).------....(((((...........))))))))).----- ( -20.80, z-score = -0.99, R) >droYak2.chrX 9121383 93 + 21770863 ----CCCACCC-CCUCCUUCAGCCACUUAUCGCUUCUUUGUGAGCGAUUGA---AUUCGAG--------UAUGUGCGCC------GCAUGUGUCGUCAUCUUAUAGAUACAUAAU----- ----..(((..-.(((.((((.......(((((((......))))))))))---)...)))--------...)))....------..(((((((...........)))))))...----- ( -17.51, z-score = -0.93, R) >droAna3.scaffold_12929 42382 115 + 3277472 CCAUACCACCCACCCCACCCCUCCCAGGGCCACUUUCUUCCAAGAUCUUGAGAUACUCAAGUAUCUAAGUGUGUGUGGCAUGUGUGUGUGUGUCGUCAUCUUGUAGAUACACAAA----- .(((((...(((((.((((((.....))).............((((((((((...)))))).))))..))).).))))...)))))((((((((...........))))))))..----- ( -34.40, z-score = -2.89, R) >droWil1.scaffold_180702 4158179 82 + 4511350 ---------------------UACAUACAUAUAUUCUAUAUGAUUUCACAAUGAAUUAAAA------AGUUCUCUCAUU------CUAUGUGUCGGCAUCUUGUAGAUAUCGAAA----- ---------------------((((((((((((...))))))..........(((((....------))))).......------.))))))(((..(((.....)))..)))..----- ( -12.50, z-score = -0.86, R) >consensus ____CCCACCC_CCUCGCCCCGCCACUUAUCGCUUCUUUGUGAGCGAUUGA___AUUCGAG________UAUGUGCGCC______GCAUGUGUCGUCAUCUUAUAGAUACAGAAU_____ ............................(((((((......)))))))........................................((((((...........))))))......... ( -7.15 = -7.36 + 0.21)

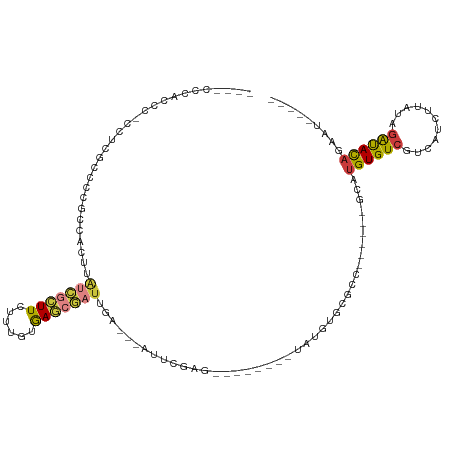

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:45:04 2011