
| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,793,590 – 14,793,756 |
| Length | 166 |
| Max. P | 0.937984 |

| Location | 14,793,590 – 14,793,690 |
|---|---|
| Length | 100 |
| Sequences | 7 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 70.51 |
| Shannon entropy | 0.54240 |
| G+C content | 0.50279 |
| Mean single sequence MFE | -29.60 |
| Consensus MFE | -8.84 |
| Energy contribution | -10.70 |
| Covariance contribution | 1.86 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.30 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.682074 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14793590 100 + 22422827 GCGUCACGGCGGACAAUUUCCGGCACUUACCACUUUUCAUACGAGUACUACCUAUGUACUAGACGC-------------AAUAUUUCUUUUGGGCCAAUUGCCCAAUCAACAA (((((..(.((((.....)))).)...................(((((.......))))).)))))-------------..........((((((.....))))))....... ( -29.20, z-score = -4.63, R) >droSim1.chrX 11424577 113 + 17042790 GCGUCACGGCGGACAAUUUCCGGCACUAACCACUUUUCAUAUGAGUACUACCUAUGUACUAGACGCCCAUGCCAGACGCAAUACAUCUUUUGGGCCAAUUGCCCAAUCAACAA (((((..((((((.....)))(((............((....))((((.......)))).....)))...))).)))))..........((((((.....))))))....... ( -32.00, z-score = -3.56, R) >droSec1.super_32 233953 113 + 562834 GCGUCACAGCGGACAAUUUCCGGCACUAACCACUUUUCAUAUGAGUACUACCUAUGUACUAGACGCCCAUGCCAGACGCAAUACAUCUUUUGGGCCAAUUGCCCAAUCAACAA (((((.....(((.....)))((((...........((....))((((.......))))..........)))).)))))..........((((((.....))))))....... ( -29.00, z-score = -3.10, R) >droYak2.chrX 9005377 113 + 21770863 GCGUCACGGCGUACAAUUUCCGGCACUUACCACUUUUCAUAUGAGCACUACCUAUGUACUAGACGCCCAUGCCAGACGCAAUAUUUCUUUUGGGCCAAUUGCCCAAUCAACAA (((((..((((((((......((......))....(((....))).........)))))..(.....)..))).)))))..........((((((.....))))))....... ( -25.70, z-score = -1.84, R) >droEre2.scaffold_4690 6541925 113 - 18748788 GCGUCACGGCGGACAAUUUCCGGCACUUACCACUUUUCAUACAAGUACUACCUAUGUGCUAGACGCCCAUGCCAGACGCAAUAUUUCUUUUGGGCCAAUUGCCCAAUCAACAA (((((..((((((.....)))(((.......((((.......)))).(((.(.....).)))..)))...))).)))))..........((((((.....))))))....... ( -29.90, z-score = -2.69, R) >dp4.chrXL_group1a 5839474 89 + 9151740 GCCUUAUGGCCGACAGUUUCCGGCACUUACGGGGACCCAUGAUGGAACUGG----GGGCUGGAUG---------------GUGGGGGCUUGCG-----UUGCCCAAUCAACAA (((....)))...((((..((((..((((.((....)).))).)...))))----..))))..((---------------((..((((.....-----..)))).)))).... ( -32.10, z-score = -0.06, R) >droPer1.super_63 267939 81 - 372969 GCCUUAUGGCCGACAGUUUCCGGCACUUACAGAGUCCCCUGAUGGAACGGG----GGAG-----------------------GGGGGCUUGGG-----UUGCCCAAUUAACAA (((((...((((........))))..........(((((((......))))----))).-----------------------.)))))(((((-----...)))))....... ( -29.30, z-score = -0.19, R) >consensus GCGUCACGGCGGACAAUUUCCGGCACUUACCACUUUUCAUAUGAGUACUACCUAUGUACUAGACGCCCAUGCCAGACGCAAUAUUUCUUUUGGGCCAAUUGCCCAAUCAACAA (((((..(.((((.....)))).)...........((......))................))))).......................((((((.....))))))....... ( -8.84 = -10.70 + 1.86)


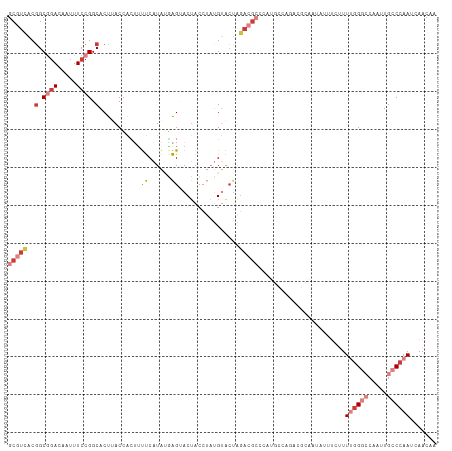
| Location | 14,793,628 – 14,793,729 |
|---|---|
| Length | 101 |
| Sequences | 8 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 69.84 |
| Shannon entropy | 0.61112 |
| G+C content | 0.40640 |
| Mean single sequence MFE | -25.16 |
| Consensus MFE | -12.80 |
| Energy contribution | -13.36 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.937984 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14793628 101 - 22422827 UCUAUUUUCACUUUUAAUUGCCCUCGAGCA-AAAAUUUGUUUGUUGAUUGGGCAAUUGGCCCAAAAGAAAUAUU--------------GCGUCUAGUACAUA-GGUAGUACUCGUAU ..((((((.....((((((((((.((((((-(....)))))))......)))))))))).......)))))).(--------------(((..((.(((...-.))).))..)))). ( -23.70, z-score = -1.79, R) >droSim1.chrX 11424615 114 - 17042790 UCUAUUUUCACUUUUAAUUGCCCUCGAGCU-AAAAUUUGUUUGUUGAUUGGGCAAUUGGCCCAAAAGAUGUAUUGCGU-CUGGCAUGGGCGUCUAGUACAUA-GGUAGUACUCAUAU ..............(((((((((.(((((.-.......)))))......)))))))))(((((..(((((.....)))-))....)))))....(((((...-....)))))..... ( -33.00, z-score = -2.24, R) >droSec1.super_32 233991 114 - 562834 UCUAUUUUCACUUUUAAUUGCCCUCGAGCU-AAAAUUUGUUUGUUGAUUGGGCAAUUGGCCCAAAAGAUGUAUUGCGU-CUGGCAUGGGCGUCUAGUACAUA-GGUAGUACUCAUAU ..............(((((((((.(((((.-.......)))))......)))))))))(((((..(((((.....)))-))....)))))....(((((...-....)))))..... ( -33.00, z-score = -2.24, R) >droYak2.chrX 9005415 114 - 21770863 UCUAUUUUCACUUUUAAUUGCCCUCGAGCU-AAAAUUUGUUUGUUGAUUGGGCAAUUGGCCCAAAAGAAAUAUUGCGU-CUGGCAUGGGCGUCUAGUACAUA-GGUAGUGCUCAUAU ..............(((((((((.(((((.-.......)))))......)))))))))(((((..(....)..(((..-...))))))))....(((((...-....)))))..... ( -28.20, z-score = -0.62, R) >droEre2.scaffold_4690 6541963 114 + 18748788 UCUAUUUUCACUUUUAAUUGCCCUCGAGCA-UAAAUUUGUUUGUUGAUUGGGCAAUUGGCCCAAAAGAAAUAUUGCGU-CUGGCAUGGGCGUCUAGCACAUA-GGUAGUACUUGUAU ..............(((((((((.((((((-......))))))......)))))))))(((((..(....)..(((..-...))))))))(((((.(.....-).))).))...... ( -29.00, z-score = -0.79, R) >droAna3.scaffold_13335 668438 115 - 3335858 UCUAUUUUCACUUUUAAUUGCCCUCGAGCC--AAGUGUGUUUGCUUAUUGGGCAAUUGGCCCACAAGAAAUAUGGCCCACAUAUUCAGGUACUAUAUACAUAUGUUAGUACCCAUAG .............((((((((((..((((.--(((....)))))))...)))))))))).......(((.((((.....))))))).((((((((((....))).)))))))..... ( -27.60, z-score = -1.46, R) >dp4.chrXL_group1a 5839512 90 - 9151740 ACUAUUUUCACUUUUAAUUGCCCGCGAGCU-AAAAUUUGUUUGUUGAUUGGGCAACGCAAGCCCCCACCAUCCAGCCC-CCAGUUCCAUCAU------------------------- .......................(.(((((-.....(((..((.((...((((.......)))).)).))..)))...-..)))))).....------------------------- ( -14.10, z-score = -0.38, R) >droPer1.super_63 267977 83 + 372969 ACUAUUUUCACUUUUAAUUGCCCGCGAGCUAAAAAUUUGUUUGUUAAUUGGGCAACCCAAGCCCCCCUC--------C-CCCGUUCCAUCAG------------------------- .................((((((((((((.........)))))).....))))))..............--------.-.............------------------------- ( -12.70, z-score = -1.47, R) >consensus UCUAUUUUCACUUUUAAUUGCCCUCGAGCU_AAAAUUUGUUUGUUGAUUGGGCAAUUGGCCCAAAAGAAAUAUUGCGU_CUGGCAUGGGCGUCUAGUACAUA_GGUAGUACUCAUAU .............((((((((((.(((((.........)))))......)))))))))).......................................................... (-12.80 = -13.36 + 0.56)


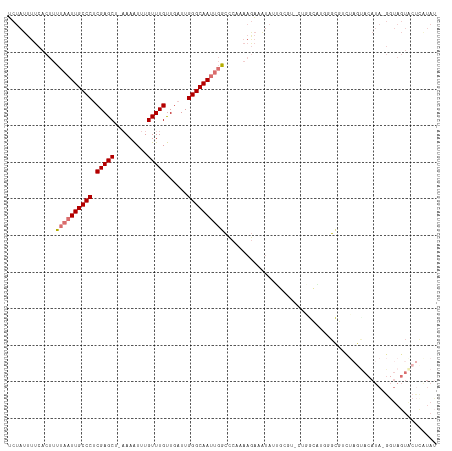
| Location | 14,793,650 – 14,793,756 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 86.56 |
| Shannon entropy | 0.25678 |
| G+C content | 0.35960 |
| Mean single sequence MFE | -26.03 |
| Consensus MFE | -19.82 |
| Energy contribution | -20.71 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.908955 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14793650 106 - 22422827 CCAACUUCAUUU--------UUUUUUGGAGCCUCUUCUAUUUUCACUUUUAAUUGCCCUCGAGCAAAAAUUUGUUUGUUGAUUGGGCAAUUGGCCCAAAAGAAAUAUUGCGUCU ..........((--------((((((((.(((..................((((((((.(((((((....)))))))......))))))))))))))))))))).......... ( -27.46, z-score = -2.77, R) >droSim1.chrX 11424650 114 - 17042790 CCAACUUCAUUUUAUUAUUUUUUUUUGGAGCCUCUUCUAUUUUCACUUUUAAUUGCCCUCGAGCUAAAAUUUGUUUGUUGAUUGGGCAAUUGGCCCAAAAGAUGUAUUGCGUCU ......................((((((.(((..................((((((((.(((((........)))))......)))))))))))))))))((((.....)))). ( -25.86, z-score = -2.41, R) >droSec1.super_32 234026 114 - 562834 CCAACUUCAUUUUAUUAUUUUUUUUUUGAGCCUCUUCUAUUUUCACUUUUAAUUGCCCUCGAGCUAAAAUUUGUUUGUUGAUUGGGCAAUUGGCCCAAAAGAUGUAUUGCGUCU .......................(((((.(((..................((((((((.(((((........)))))......))))))))))).)))))((((.....)))). ( -22.76, z-score = -1.81, R) >droYak2.chrX 9005450 113 - 21770863 CCAACUUUAUUUUCAUA-UAUUUUUUGGAGCCUCUUCUAUUUUCACUUUUAAUUGCCCUCGAGCUAAAAUUUGUUUGUUGAUUGGGCAAUUGGCCCAAAAGAAAUAUUGCGUCU ..............(((-(.((((((((.(((..................((((((((.(((((........)))))......))))))))))))))))))).))))....... ( -25.76, z-score = -2.35, R) >droEre2.scaffold_4690 6541998 113 + 18748788 CCAACUUUGCUUUAUU-UAUUUGUUCGGAGCCUCUUCUAUUUUCACUUUUAAUUGCCCUCGAGCAUAAAUUUGUUUGUUGAUUGGGCAAUUGGCCCAAAAGAAAUAUUGCGUCU ........((..((((-(.(((....((.(((..................((((((((.((((((......))))))......))))))))))))).))).)))))..)).... ( -25.16, z-score = -1.66, R) >droAna3.scaffold_13335 668478 106 - 3335858 CCAACUUUUAUU----AUUACUUGGCAGGGCCUCUUCUAUUUUCACUUUUAAUUGCCCUCGAGCC-AAGUGUGUUUGCUUAUUGGGCAAUUGGCCCACAAGAAAUAUGGCC--- (((.........----..(((((((((((((.......................)))))...)))-)))))(((((.(((..(((((.....))))).)))))))))))..--- ( -29.20, z-score = -1.62, R) >consensus CCAACUUCAUUUUAUUAUUUUUUUUUGGAGCCUCUUCUAUUUUCACUUUUAAUUGCCCUCGAGCUAAAAUUUGUUUGUUGAUUGGGCAAUUGGCCCAAAAGAAAUAUUGCGUCU ....................((((((((.(((..................((((((((.(((((........)))))......)))))))))))))))))))............ (-19.82 = -20.71 + 0.89)


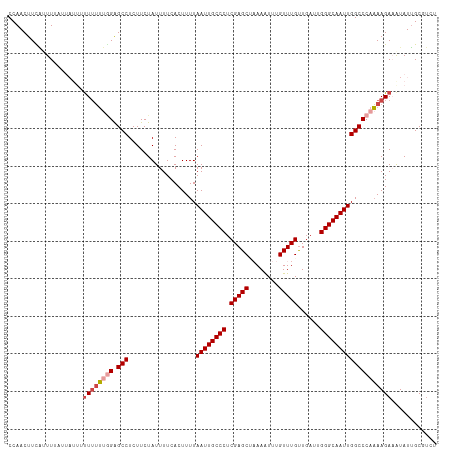
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:44:48 2011