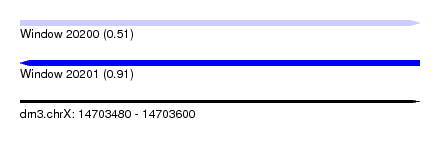

| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,703,480 – 14,703,600 |
| Length | 120 |
| Max. P | 0.905666 |

| Location | 14,703,480 – 14,703,600 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.03 |
| Shannon entropy | 0.19671 |
| G+C content | 0.43882 |
| Mean single sequence MFE | -32.78 |
| Consensus MFE | -26.01 |
| Energy contribution | -27.85 |
| Covariance contribution | 1.84 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.506500 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14703480 120 + 22422827 AAGGACCUGCCCAAGUCGUCCCAGGAUGUGGGUAUGAGUCCAAUGUCCUCUUUAGACCAAUGUUGUUUACGACAUUUAUUGUGACUGCAAAACCACACGAAUUAAAGGACUUGGAUAAUA ..(((((((((((...((((....)))))))))).).))))..((((((((((((...(((((((....)))))))..(((((............))))).)))))))....)))))... ( -33.70, z-score = -1.76, R) >droSim1.chrX 11354111 119 + 17042790 GAGGACCUGCCCAAGUCCUCCCAGGAUGUGGGUAUGAGUCCAAUGUCCUCUUUAGACCAAUGUUGUUUACGACACUUAUUGUGACUGCAGA-CCACACGAAUUAAAGGACUUGGAUAAUA ..........(((((((((...((..(((((((.(((((((((((((.......)))...(((((....)))))...)))).)))).)).)-)).))))..))..)))))))))...... ( -35.10, z-score = -1.85, R) >droSec1.super_32 150325 116 + 562834 GAGGACCUGCCCAAGUCCUCCCAGGAUGUGGGUAUGAGUCCAAUGUCCUCUUUAGACCCAUGUUGUUUACGACACUUAUUGUGACUGCAGA-CCACACGAAUUAAAGGACUUGGAUA--- ..........(((((((((...((..(((((((.(((((((((((((.......)))...(((((....)))))...)))).)))).)).)-)).))))..))..)))))))))...--- ( -35.10, z-score = -1.79, R) >droYak2.chrX 8924091 108 + 21770863 -----------AAGGACCUCCCAGGAUGUGGGUAUGAGUCCAAUGUCCUCUUUGGACCAAUGUUGUUUACGACAUUUAUUGUGACUGCAGA-CCACUCGAAUUAAAGGACUGGAAUAAUA -----------.........((((...((((...((((((((((((((.....)))).(((((((....))))))).)))).)))).))..-))))((........)).))))....... ( -29.70, z-score = -1.12, R) >droEre2.scaffold_4690 6458155 118 - 18748788 AAGGACCUGCC-AAGUCCUUCCAGGAUGUGGGUAUGAGUCCAAUGUCCUUUUUAGACCAAUGUUGCUUACGACAUUUAUUGUGACUGCAGA-CCACUCGAAUUAAAUGACUUGGAUAAUA .(((((.....-..)))))((((((..((((...(((((((((((((.......))).(((((((....))))))).)))).)))).))..-))))(((.......)))))))))..... ( -30.30, z-score = -1.34, R) >consensus AAGGACCUGCCCAAGUCCUCCCAGGAUGUGGGUAUGAGUCCAAUGUCCUCUUUAGACCAAUGUUGUUUACGACAUUUAUUGUGACUGCAGA_CCACACGAAUUAAAGGACUUGGAUAAUA ..........(((((((((........((((...(((((((((((((.......))).(((((((....))))))).)))).)))).))...)))).........)))))))))...... (-26.01 = -27.85 + 1.84)

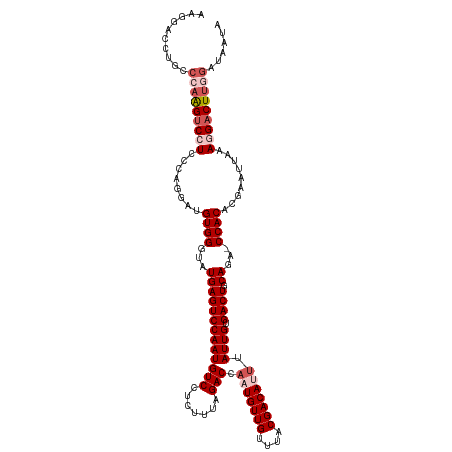
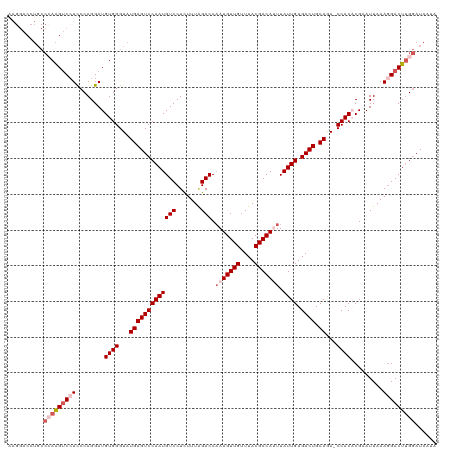
| Location | 14,703,480 – 14,703,600 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.03 |
| Shannon entropy | 0.19671 |
| G+C content | 0.43882 |
| Mean single sequence MFE | -33.70 |
| Consensus MFE | -30.26 |
| Energy contribution | -32.10 |
| Covariance contribution | 1.84 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.90 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.18 |
| SVM RNA-class probability | 0.905666 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14703480 120 - 22422827 UAUUAUCCAAGUCCUUUAAUUCGUGUGGUUUUGCAGUCACAAUAAAUGUCGUAAACAACAUUGGUCUAAAGAGGACAUUGGACUCAUACCCACAUCCUGGGACGACUUGGGCAGGUCCUU .....((((((((.(((.....((((((...((.((((.((((.(((((.(....).))))).((((.....))))))))))))))...))))))....))).))))))))......... ( -33.60, z-score = -1.57, R) >droSim1.chrX 11354111 119 - 17042790 UAUUAUCCAAGUCCUUUAAUUCGUGUGG-UCUGCAGUCACAAUAAGUGUCGUAAACAACAUUGGUCUAAAGAGGACAUUGGACUCAUACCCACAUCCUGGGAGGACUUGGGCAGGUCCUC .....((((((((((((.....((((((-..((.((((.((((.(((((.(....).))))).((((.....))))))))))))))...))))))....))))))))))))......... ( -39.10, z-score = -2.53, R) >droSec1.super_32 150325 116 - 562834 ---UAUCCAAGUCCUUUAAUUCGUGUGG-UCUGCAGUCACAAUAAGUGUCGUAAACAACAUGGGUCUAAAGAGGACAUUGGACUCAUACCCACAUCCUGGGAGGACUUGGGCAGGUCCUC ---..((((((((((((.((((((((.(-(.(((.(.(((.....))).)))).)).))))))))....))))))).)))))......((((.....))))(((((((....))))))). ( -40.60, z-score = -2.72, R) >droYak2.chrX 8924091 108 - 21770863 UAUUAUUCCAGUCCUUUAAUUCGAGUGG-UCUGCAGUCACAAUAAAUGUCGUAAACAACAUUGGUCCAAAGAGGACAUUGGACUCAUACCCACAUCCUGGGAGGUCCUU----------- ..........(.(((((.....((((((-..((.((((.((((.(((((.(....).))))).((((.....))))))))))))))...)))).))...))))).)...----------- ( -25.70, z-score = -0.53, R) >droEre2.scaffold_4690 6458155 118 + 18748788 UAUUAUCCAAGUCAUUUAAUUCGAGUGG-UCUGCAGUCACAAUAAAUGUCGUAAGCAACAUUGGUCUAAAAAGGACAUUGGACUCAUACCCACAUCCUGGAAGGACUU-GGCAGGUCCUU ......(((((((.(((.....((((((-..((.((((.((((.(((((.(....).))))).((((.....))))))))))))))...)))).))...))).)))))-))......... ( -29.50, z-score = -1.17, R) >consensus UAUUAUCCAAGUCCUUUAAUUCGUGUGG_UCUGCAGUCACAAUAAAUGUCGUAAACAACAUUGGUCUAAAGAGGACAUUGGACUCAUACCCACAUCCUGGGAGGACUUGGGCAGGUCCUC .....((((((((((((.....((((((...((.((((.((((.(((((.(....).))))).((((.....))))))))))))))...))))))....))))))))))))......... (-30.26 = -32.10 + 1.84)

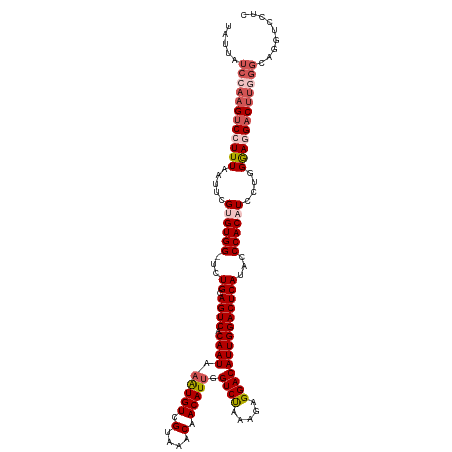
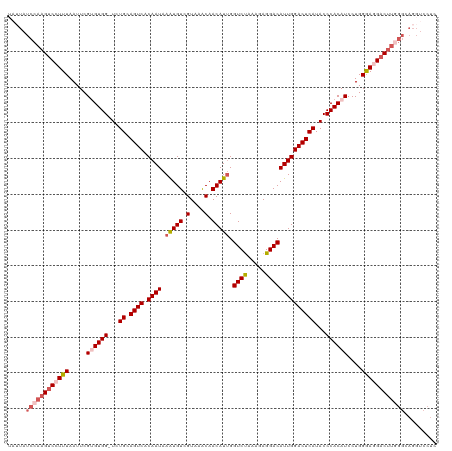
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:44:34 2011