
| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,655,878 – 14,655,996 |
| Length | 118 |
| Max. P | 0.513035 |

| Location | 14,655,878 – 14,655,973 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 63.71 |
| Shannon entropy | 0.70674 |
| G+C content | 0.32381 |
| Mean single sequence MFE | -14.18 |
| Consensus MFE | -5.70 |
| Energy contribution | -6.45 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.53 |
| Mean z-score | -0.86 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.513035 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
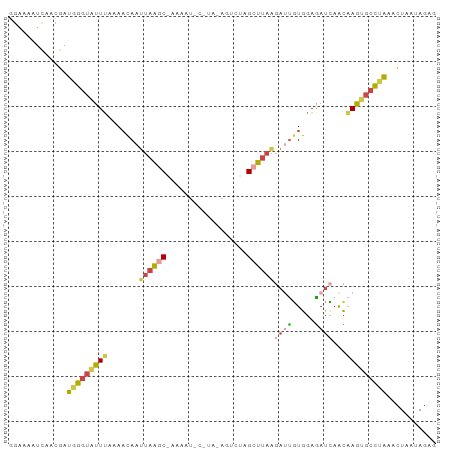
>dm3.chrX 14655878 95 + 22422827 GGAAAAUCAACGAUGGGUAUUUAAAACAAUUAAGCAAAAAUACAUAGAGUCUAGCUUAUGAUUGUGGAGAUCAACAAGUGCCUAAACUAAUAGAG .............(((((((((........(((((..................)))))(((((.....)))))..)))))))))........... ( -14.37, z-score = -0.59, R) >droSim1.chrX 11314206 81 + 17042790 GGAAAAUCAACGAUGGGUAUUUAAAACAAUUAAGC--------------UUUAGCUUAAGAUUGUGGAGAUCAACAAGUGCCUAAACUAAUAGAG .............(((((((((.......((((((--------------....))))))((((.....))))...)))))))))........... ( -16.50, z-score = -2.20, R) >droSec1.super_32 108146 81 + 562834 GGAAAAUCAACGAUGGGUAUUUAAAACAAUUAAGC--------------UUUAGCUUAAGAUUGUGGAGAUCAACAAGUGCCUAAACUAAUAGAG .............(((((((((.......((((((--------------....))))))((((.....))))...)))))))))........... ( -16.50, z-score = -2.20, R) >droYak2.chrX 8882324 95 + 21770863 CGUAAAUCAACGAUGGGUAUUUAAAACAAUUAAACGAAAAUACAUAAAGUCUAGCUCAAGAUUGUGGAGAUCAACAAGUGCCUAAACUAACAGAA (((......))).(((((((((.........................(((((......)))))((........)))))))))))........... ( -11.90, z-score = 0.02, R) >droEre2.scaffold_4690 6418774 82 - 18748788 ----------CGUUGGUUGUUCAAAACAAUUAAACAAAAAUACUUAAACAAUAGCUUAG---CUUGGAGAUAAACUAAUGCCCAAACUAAUAGAG ----------..((((((((.....))))))))......................((((---.((((..((......))..)))).))))..... ( -8.90, z-score = 0.08, R) >droGri2.scaffold_15245 3897516 91 - 18325388 UAAAUGUUACAAGAAAGUAUUGCCACUAAUUAGGCGCCAGU-UGAAGAGCGCAGCUUUA-ACUGUAUAAAUUUGUUAA--CUCUAGUUUAUAUUU .((((((.((.(((.......(((........)))..((((-(((((.......)))))-))))..............--.))).))..)))))) ( -16.90, z-score = -0.27, R) >consensus GGAAAAUCAACGAUGGGUAUUUAAAACAAUUAAGC_AAAAU_C_UA_AGUCUAGCUUAAGAUUGUGGAGAUCAACAAGUGCCUAAACUAAUAGAG .............(((((((((.......((((((..................))))))((((.....))))...)))))))))........... ( -5.70 = -6.45 + 0.76)


| Location | 14,655,896 – 14,655,996 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 65.59 |
| Shannon entropy | 0.65409 |
| G+C content | 0.32947 |
| Mean single sequence MFE | -20.30 |
| Consensus MFE | -5.44 |
| Energy contribution | -6.64 |
| Covariance contribution | 1.20 |
| Combinations/Pair | 1.42 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.27 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.507416 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
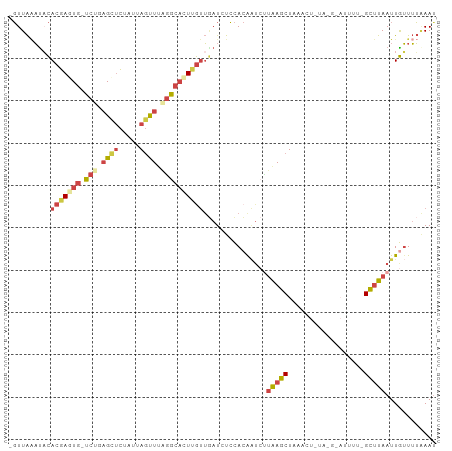
>dm3.chrX 14655896 100 - 22422827 -GUUAAAUACACGAGUGAUCUGAGCUCUAUUAGUUUAGGCACUUGUUGAUCUCCACAAUCAUAAGCUAGACUCUAUGUAUUUUUGCUUAAUUGUUUUAAAU -.(((((...(((((((.((((((((.....)))))))))))))))........(((((..(((((...((.....))......))))))))))))))).. ( -21.50, z-score = -1.70, R) >droSim1.chrX 11314224 85 - 17042790 -GUUAAAUACACGAGUG-UCUGAGCUCUAUUAGUUUAGGCACUUGUUGAUCUCCACAAUCUUAAGCUAAA--------------GCUUAAUUGUUUUAAAU -.(((((...(((((((-((((((((.....)))))))))))))))........(((((..(((((....--------------))))))))))))))).. ( -25.60, z-score = -4.47, R) >droSec1.super_32 108164 85 - 562834 -GUUAAACACACGAGUG-UCUGAGCUCUAUUAGUUUAGGCACUUGUUGAUCUCCACAAUCUUAAGCUAAA--------------GCUUAAUUGUUUUAAAU -.(((((((.(((((((-((((((((.....)))))))))))))))))......(((((..(((((....--------------))))))))))))))).. ( -26.40, z-score = -4.65, R) >droYak2.chrX 8882342 99 - 21770863 -GUUAAAUACACGAAUG-CCGCUGUUCUGUUAGUUUAGGCACUUGUUGAUCUCCACAAUCUUGAGCUAGACUUUAUGUAUUUUCGUUUAAUUGUUUUAAAU -...(((((((.((((.-.....))))..((((((((((...((((.(....).)))).))))))))))......)))))))..((((((.....)))))) ( -16.00, z-score = -0.13, R) >droEre2.scaffold_4690 6418782 96 + 18748788 -GUUACAUUCACGAAUG-CCGCAGCUCUAUUAGUUUGGGCAUUAGUUUAUCUCCAAG---CUAAGCUAUUGUUUAAGUAUUUUUGUUUAAUUGUUUUGAAC -...........(((((-(.((((.....((((((((((.((......)).))))))---))))....))))....))))))..((((((.....)))))) ( -16.50, z-score = -0.20, R) >droGri2.scaffold_15245 3897534 97 + 18325388 GUUUGGUUCUAAUACUUAUAUAAGAAAUAUAAACUAGAG--UUAACAAAUUUAUACAGU-UAAAGCUGCGCUCUUCA-ACUGGCGCCUAAUUAGUGGCAAU (((((.((((.(((....))).))))...)))))(((..--(((((...........))-)))..))).(((.....-...)))(((........)))... ( -15.80, z-score = 0.13, R) >consensus _GUUAAAUACACGAGUG_UCUGAGCUCUAUUAGUUUAGGCACUUGUUGAUCUCCACAAUCUUAAGCUAAACU_UA_G_AUUUU_GCUUAAUUGUUUUAAAU ..........(((((((.(((.((((.....)))).))))))))))...............(((((..................)))))............ ( -5.44 = -6.64 + 1.20)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:44:24 2011