
| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,621,918 – 14,622,050 |
| Length | 132 |
| Max. P | 0.548765 |

| Location | 14,621,918 – 14,622,050 |
|---|---|
| Length | 132 |
| Sequences | 6 |
| Columns | 140 |
| Reading direction | reverse |
| Mean pairwise identity | 75.25 |
| Shannon entropy | 0.46655 |
| G+C content | 0.55447 |
| Mean single sequence MFE | -33.11 |
| Consensus MFE | -19.22 |
| Energy contribution | -20.83 |
| Covariance contribution | 1.61 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.548765 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14621918 132 - 22422827 CCCAGGCUGUUUAUACGCCCCUCCACUCCAU-----CGCCACUGUCCUCGAUUUGCCCCCUUAUCUGCAGC---GCACUGUAUCUGUGCCGCAAAUCACAGUCUGUGGGGCGAACGCGUCGUAUACGUGUUAUACCUACG ...(((((((......((.............-----.))..........(((((((.....((..(((((.---...)))))..))....))))))))))))))(((((...(((((((.....)))))))...))))). ( -34.54, z-score = -0.95, R) >droSim1.chrX 11281821 132 - 17042790 CCCAGGCUGUUUAUACGCCCCUCCACUCCAU-----CCCCACUGCCCUCGAUUUGCCUCCUUAUCUGCAGC---GCACUGUAUCUGUGCCGCAAAUCACAGUCUGUGGGGCGAACGCGUCGUAUACGUGUUAUACCUACG ...(((((((......((.............-----.......))....(((((((((.(......).)).---((((.......)))).))))))))))))))(((((...(((((((.....)))))))...))))). ( -34.65, z-score = -1.46, R) >droSec1.super_32 75364 132 - 562834 CCCAGGCUGUUUAUACGCCCCUCCACUCCAU-----CCCCACUGCCCUCGAUUUGCCCCCUUAUCUGCAGC---GCACUGUAUCUGUGCCGCAAAUCACAGUCUGUGGGGCGAACGCGUCGUAUACGUGUUAUACCUACG ...(((((((......((.............-----.......))....(((((((.....((..(((((.---...)))))..))....))))))))))))))(((((...(((((((.....)))))))...))))). ( -34.05, z-score = -1.27, R) >droYak2.chrX 8848602 137 - 21770863 CCCAUGCUGUUUAUACGCCCCUCCACUCCACUCCUGCCCCAAUGUCCUCGAUAUGCCUCCUUAUCUGCAGC---CCACUGUAUCUGUGCCGCAAAUCACAGUCUGUGGGGCGAACGCGUCGUAUACGUGUUAUACCAACG ........(((....((((((............................((((........)))).((((.---..(((((((.((.....)).)).)))))))))))))))(((((((.....))))))).....))). ( -30.00, z-score = -0.80, R) >droEre2.scaffold_4690 6383622 132 + 18748788 CCCGUGCUGUUUAUACGCCCCUCUACGCCUG-----CCCCAAUGUUCUCGAUAUACCUCCUUAUCUGCAGC---CCACUGUAUCUGUGCCGCAAAUCACAGUCUGUGGGGCGAACGCGUCGUAUACGUGUUGUACCUACG ...((((.....(((((.....(.((((.((-----(((((........((((........))))(((((.---...))))).(((((........)))))....)))))))...)))).)....))))).))))..... ( -33.40, z-score = -1.19, R) >droAna3.scaffold_12948 24821 101 + 692136 -------------------------------------CUUAUUUUGUUUAUGUGGGAGCGCCGUCUGCUCCCUUCCAGUAUCUCCGGGUGGCAAAUCACGGUCGCCAGAGCCCACCCGGCGUAUAC--GUUAUGCUUAUG -------------------------------------................(((((((.....))))))).....(((((.((((((((........(.((....)).))))))))).).))))--............ ( -32.00, z-score = -1.31, R) >consensus CCCAGGCUGUUUAUACGCCCCUCCACUCCAU_____CCCCACUGUCCUCGAUUUGCCUCCUUAUCUGCAGC___CCACUGUAUCUGUGCCGCAAAUCACAGUCUGUGGGGCGAACGCGUCGUAUACGUGUUAUACCUACG ...............((((((............................................(((((.......))))).(((((........))))).....))))))((((((.......))))))......... (-19.22 = -20.83 + 1.61)

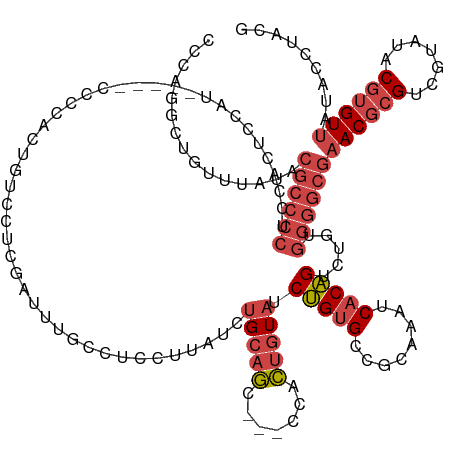
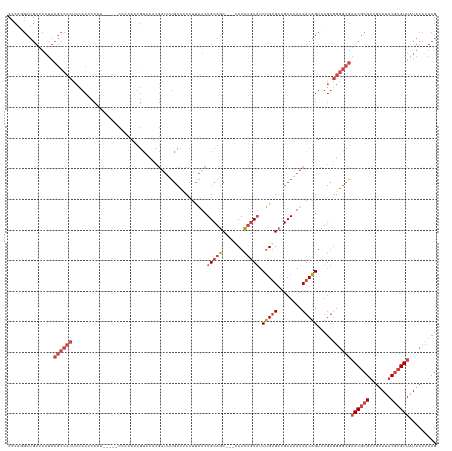
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:44:15 2011