| Sequence ID | dm3.chr2L |
|---|---|
| Location | 11,400,069 – 11,400,169 |
| Length | 100 |
| Max. P | 0.955905 |

| Location | 11,400,069 – 11,400,169 |
|---|---|
| Length | 100 |
| Sequences | 11 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 92.73 |
| Shannon entropy | 0.15723 |
| G+C content | 0.37872 |
| Mean single sequence MFE | -25.93 |
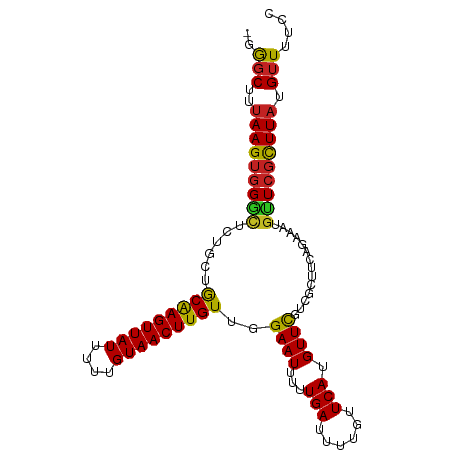
| Consensus MFE | -20.83 |
| Energy contribution | -20.39 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.29 |
| Mean z-score | -2.60 |
| Structure conservation index | 0.80 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.62 |
| SVM RNA-class probability | 0.955905 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
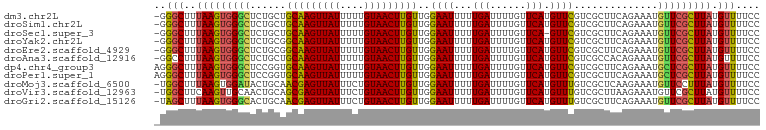
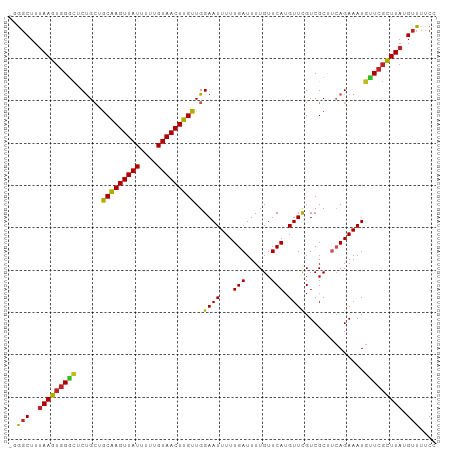
>dm3.chr2L 11400069 100 + 23011544 -GGGCUUUAAGUGGGCUCUGCUGCAAGUUAUUUUUGUAACUUGUUGGAAUUUUUGAUUUUGUUCAUGUUCGUCGCUUCAGAAAUGUUCGCUUAUGUUUUCC -((((..(((((((((((((..(((((((((....)))))))....((((...(((......))).))))...))..))))...))))))))).))))... ( -25.60, z-score = -2.70, R) >droSim1.chr2L 11208436 100 + 22036055 -GGGCUUUAAGUGGGCUCUGCUGCAAGUUAUUUUUGUAACUUGUUGGAAUUUUUGAUUUUGUUCAUGUUCGUCGCUUCAGAAAUGUUCGCUUAUGUUUUCC -((((..(((((((((((((..(((((((((....)))))))....((((...(((......))).))))...))..))))...))))))))).))))... ( -25.60, z-score = -2.70, R) >droSec1.super_3 6794828 99 + 7220098 -GGGCUUUAAGUGGGCUCUGCUGCAAGUUAUUUUUGUAACUUGUUGGAAUUUUUGAUUUUGUUCA-GUUCGUCGCUUCAGAAAUGUUCGCUUAUGUUUUCC -((((..(((((((((((((..(((((((((....)))))))....((....((((......)))-)....))))..))))...))))))))).))))... ( -24.70, z-score = -2.22, R) >droYak2.chr2L 7798328 100 + 22324452 -GGGCUUUAAGUGGGCUCUGCGGCAAGUUAUUUUUGUAACUUGUUGGAAUUUUUGAUUUUGUUCAUGUUCGUCGCUUCAGAAAUGUUCGCUUAUGUUUUCC -((((..((((((((((((((((((((((((....)))))))))))((((...(((......))).)))).......))))...))))))))).))))... ( -28.50, z-score = -3.44, R) >droEre2.scaffold_4929 12601610 100 - 26641161 -GGGCUUUAAGUGGGCUCUGCGGCAAGUUAUUUUUGUAACUUGUUGGAAUUUUUGAUUUUGUUCAUGUUCGUCGCUUCAGAAAUGUUCGCUUAUGUUUUCC -((((..((((((((((((((((((((((((....)))))))))))((((...(((......))).)))).......))))...))))))))).))))... ( -28.50, z-score = -3.44, R) >droAna3.scaffold_12916 12108970 100 - 16180835 -GGCCUUUAAGUGGGCUCUGCUGCAAGUUAUUUUUGUAACUUGUUGGAAUUUUUGAUUUUGUUCAUGUUCGUCGCCACAGAAAUGUUCGCUUAUGUUUUCC -......(((((((((....(.(((((((((....))))))))).)((((...(((......))).))))...)))(((....))).))))))........ ( -25.20, z-score = -2.23, R) >dp4.chr4_group3 6851077 101 + 11692001 AGGGCUUUAAGUGGGCUCCGGUGCAAGUUAUUUUUGUAACUUGUUGGAAUUUUUGAUUUUGUUCAUGUUCGUCGCUUCAGAAAUGCUCGCUUAUGUUUUCC (((((..(((((((((((((..(((((((((....)))))))))))))((((((((...((........)).....))))))))))))))))).))))).. ( -29.60, z-score = -3.36, R) >droPer1.super_1 8299017 101 + 10282868 AGGGCUUUAAGUGGGCUCCGGUGCAAGUUAUUUUUGUAACUUGUUGGAAUUUUUGAUUUUGUUCAUGUUCGUCGCUUCAGAAAUGCUCGCUUAUGUUUUCC (((((..(((((((((((((..(((((((((....)))))))))))))((((((((...((........)).....))))))))))))))))).))))).. ( -29.60, z-score = -3.36, R) >droMoj3.scaffold_6500 5057773 100 - 32352404 -UGGCUUUAAGUGGAUACUGCAACGAGUUAUUUCUGUAACUUGUUGGAAUUUUUGAUUUUGUUCAUGUUUGUCGCUCAAGAAAUGUUCCUUUAUGUUUUCC -.(((.....(((((((...(((((((((((....)))))))))))(((((...))))))))))))....)))......(((((((......))))))).. ( -20.00, z-score = -1.28, R) >droVir3.scaffold_12963 18075412 100 - 20206255 -UGGCUUCAAGUUGCAACUGCAGCGAGUUAUUUCUGUAACUUGUUGGAAUUUUUGAUUUUGUUCAUGUUUGUCGCUUAAGAAAUGUUCGCUUAUGUUUUCC -...(((.((((.((((...(((((((((((....)))))))))))((((..........))))....)))).)))))))..................... ( -19.90, z-score = -0.42, R) >droGri2.scaffold_15126 7217679 100 - 8399593 -UAGCUUUAAGUGGGCACUGCAACGAGUUAUUUCUGUAACUUGUUGGAAUUUUUGAUUUUGUUCAUGUUUGUCGCUUCAGAAAUGUUCGCUUAUGUUUUCC -.(((..(((((((((....(((((((((((....)))))))))))..((((((((....((.((....))..)).))))))))))))))))).))).... ( -28.00, z-score = -3.47, R) >consensus _GGGCUUUAAGUGGGCUCUGCUGCAAGUUAUUUUUGUAACUUGUUGGAAUUUUUGAUUUUGUUCAUGUUCGUCGCUUCAGAAAUGUUCGCUUAUGUUUUCC ..(((..(((((((((......(((((((((....)))))))))..((((...(((......))).))))..............))))))))).))).... (-20.83 = -20.39 + -0.44)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:32:25 2011