
| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,494,365 – 14,494,475 |
| Length | 110 |
| Max. P | 0.694144 |

| Location | 14,494,365 – 14,494,466 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 90.04 |
| Shannon entropy | 0.17968 |
| G+C content | 0.44320 |
| Mean single sequence MFE | -28.98 |
| Consensus MFE | -25.40 |
| Energy contribution | -25.64 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.88 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.539913 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
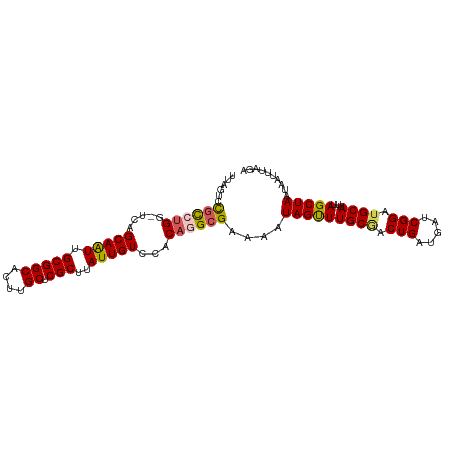
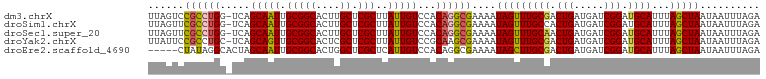
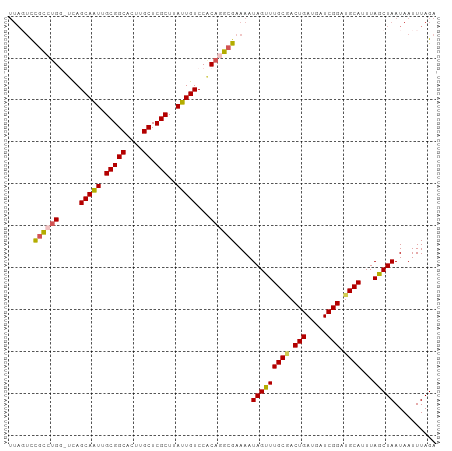
>dm3.chrX 14494365 101 - 22422827 UUAGUCCGCCUGG-UCAGCAAUUGCGGCACUUGCUCGCUUAUUGUCCACAGGCGAAAAUAGUUUGCGACUGAUGAUCGGAUGCAUUUAGCUAAUAAUUUAGA ......((((((.-...(((((.(((((....)).)))..)))))...))))))....(((((((((.(((.....))).))))...))))).......... ( -30.60, z-score = -1.61, R) >droSim1.chrX 11186131 101 - 17042790 UUAGUUCGCCUGG-UCAGCAAUUGCGGCACUUGCUCGCUUAUUGUCCACAGGCGAAAAUAGUUUGCCACUGAUGAUCGGAUGCAUUUAGCUAAUAAUUUAGA ....((((((((.-...(((((.(((((....)).)))..)))))...))))))))..((((((((..(((.....)))..)))...))))).......... ( -30.70, z-score = -1.86, R) >droSec1.super_20 1085487 101 - 1148123 UUAGUUCGCCUGG-UCAGCAAUUGCGGCACUUGCUCGCUUAUUGUCCACAGGCGAAAAUAGUUUGCAACUGAUGAUCGGAUGCAUUUAGCUAAUAAUUUAGA ....((((((((.-...(((((.(((((....)).)))..)))))...))))))))..(((((((((.(((.....))).))))...))))).......... ( -33.70, z-score = -2.91, R) >droYak2.chrX 8728601 101 - 21770863 UUAUUCCGCCUGC-UCAGCAGUUGCGGCACUCGCUCGCUUAUUGUCCGCAAGCGAAAAUAGUUUGCGACUGAUGAUCGGAUGCAUUUAGCUAAUAAUUUAGA .......(((((.-(((.(((((((((.(((...((((((.........))))))....)))))))))))).))).)))..))......((((....)))). ( -26.70, z-score = -0.11, R) >droEre2.scaffold_4690 6269516 97 + 18748788 -----CUAUAGGCACUAGCAAUUGCGGCACUGGCUCGCUCAUUGUCCACAGGCGAAAAUAGCUUGCGACUGAUGAUCGGAUGCAUUUAGCUAAUAAUUUAGA -----(((.......((((((.((((((....))(((.((((((((..(((((.......))))).))).))))).))).)))).)).))))......))). ( -23.22, z-score = 0.37, R) >consensus UUAGUCCGCCUGG_UCAGCAAUUGCGGCACUUGCUCGCUUAUUGUCCACAGGCGAAAAUAGUUUGCGACUGAUGAUCGGAUGCAUUUAGCUAAUAAUUUAGA ......((((((.....(((((.(((((....)).)))..)))))...))))))....(((((((((.(((.....))).))))...))))).......... (-25.40 = -25.64 + 0.24)

| Location | 14,494,372 – 14,494,475 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 91.84 |
| Shannon entropy | 0.15212 |
| G+C content | 0.46388 |
| Mean single sequence MFE | -32.68 |
| Consensus MFE | -25.80 |
| Energy contribution | -26.56 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.694144 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
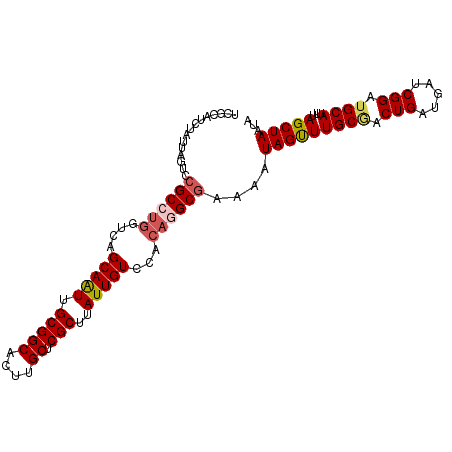
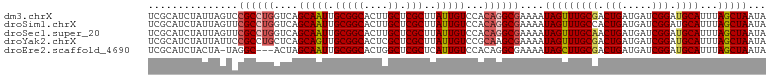
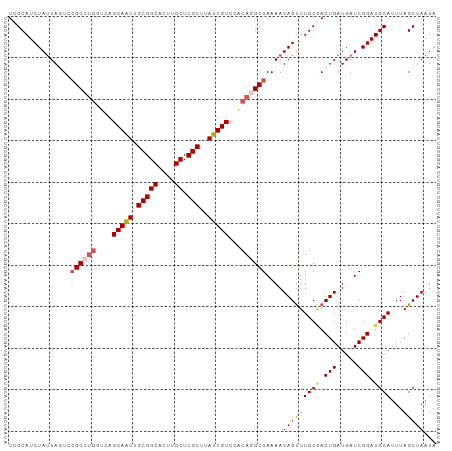
>dm3.chrX 14494372 103 - 22422827 UCGCAUCUAUUAGUCCGCCUGGUCAGCAAUUGCGGCACUUGCUCGCUUAUUGUCCACAGGCGAAAAUAGUUUGCGACUGAUGAUCGGAUGCAUUUAGCUAAUA ..(((((((((((((((((((....(((((.(((((....)).)))..)))))...))))))......(....))))))))....))))))............ ( -35.50, z-score = -2.65, R) >droSim1.chrX 11186138 103 - 17042790 UCGCAUCUAUUAGUUCGCCUGGUCAGCAAUUGCGGCACUUGCUCGCUUAUUGUCCACAGGCGAAAAUAGUUUGCCACUGAUGAUCGGAUGCAUUUAGCUAAUA ..((((((((((.((((((((....(((((.(((((....)).)))..)))))...))))))))..((((.....)))).)))).))))))............ ( -34.40, z-score = -2.62, R) >droSec1.super_20 1085494 103 - 1148123 UCGCAUCUAUUAGUUCGCCUGGUCAGCAAUUGCGGCACUUGCUCGCUUAUUGUCCACAGGCGAAAAUAGUUUGCAACUGAUGAUCGGAUGCAUUUAGCUAAUA ..((..(((((..((((((((....(((((.(((((....)).)))..)))))...)))))))))))))..((((.(((.....))).))))....))..... ( -35.10, z-score = -2.92, R) >droYak2.chrX 8728608 103 - 21770863 UCGCAUCUAUUAUUCCGCCUGCUCAGCAGUUGCGGCACUCGCUCGCUUAUUGUCCGCAAGCGAAAAUAGUUUGCGACUGAUGAUCGGAUGCAUUUAGCUAAUA ..(((((((((((.((((((((...))))..))))((.((((..(((.(((.(((....).)).))))))..)))).))))))).))))))............ ( -31.50, z-score = -1.20, R) >droEre2.scaffold_4690 6269523 99 + 18748788 UCGCAUCUACUA-UAGGC---ACUAGCAAUUGCGGCACUGGCUCGCUCAUUGUCCACAGGCGAAAAUAGCUUGCGACUGAUGAUCGGAUGCAUUUAGCUAAUA ..((((((....-...((---....))....(((((....)).)))((((((((..(((((.......))))).))).)))))..))))))............ ( -26.90, z-score = -0.41, R) >consensus UCGCAUCUAUUAGUCCGCCUGGUCAGCAAUUGCGGCACUUGCUCGCUUAUUGUCCACAGGCGAAAAUAGUUUGCGACUGAUGAUCGGAUGCAUUUAGCUAAUA ...............((((((....(((((.(((((....)).)))..)))))...))))))....(((((((((.(((.....))).))))...)))))... (-25.80 = -26.56 + 0.76)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:43:54 2011