
| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,469,033 – 14,469,147 |
| Length | 114 |
| Max. P | 0.523690 |

| Location | 14,469,033 – 14,469,147 |
|---|---|
| Length | 114 |
| Sequences | 8 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.63 |
| Shannon entropy | 0.29334 |
| G+C content | 0.37939 |
| Mean single sequence MFE | -24.45 |
| Consensus MFE | -17.86 |
| Energy contribution | -17.68 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.73 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.523690 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14469033 114 - 22422827 AUGUUAAUGCU-AUUGGU----UGCCACAGUGACCAAAAUUGAUGUCAAUUGAUAUUGAUAAUCCAUGCAUUAUGCGCGAUAAACAAAGUUAAUUGAAUCAAAGGCGCCGAGCCAACAA- .((((......-.(((((----..(....)..)))))..(((((.(((((((((.(((...(((..(((.....))).)))...))).)))))))))))))).(((.....))))))).- ( -30.30, z-score = -2.32, R) >droAna3.scaffold_13248 2190277 114 - 4840945 AUGUUAAUGCU-AUUGGU----UUCCCGAGCAACCAAAAUUGAUGUCAAUUGAUAUUGAUAAUCCAUGCAUUAUGCGCGAUAAACAAAGUUAAUUGAAUCAAAGGCGCU-ACCCUCCACU ........(((-.(((((----(........))))))..(((((.(((((((((.(((...(((..(((.....))).)))...))).)))))))))))))).)))...-.......... ( -23.00, z-score = -1.15, R) >droSim1.chrX 11149462 114 - 17042790 AUGUUAAUGCU-AUUGGU----UGCCACAGUGACCAAAAUUGAUGUCAAUUGAUAUUGAUAAUCCAUGCAUUAUGCGCGAUAAACAAAGUUAAUUGAAUCAAAGGCGCCGAGCCCACGA- ...........-.(((((----..(....)..)))))..(((((.(((((((((.(((...(((..(((.....))).)))...))).)))))))))))))).(((.....))).....- ( -28.10, z-score = -1.50, R) >droSec1.super_20 1060846 114 - 1148123 AUGUUAAUGCU-AUUGGU----UGCCACAGUGACCAAAAUUGAUGUCAAUUGAUAUUGAUAAUCCAUGCAUUAUGCGCGAUAAACAAAGUUAAUUGAAUCAAAGGCGCCGAGCCCACAA- ...........-.(((((----..(....)..)))))..(((((.(((((((((.(((...(((..(((.....))).)))...))).)))))))))))))).(((.....))).....- ( -28.10, z-score = -1.62, R) >droYak2.chrX 8704315 114 - 21770863 UUGUUAAUGCU-AUUGGU----UUCCACAGUGACCGAAAUUGAUGUCAAUUGAUAUUGAUAAUCCAUGCAUUAUGCGCGAUAAACAAAGUUAAUUGAAUCAAAGGCCCUGAACCCACAA- ..(((...(((-.(((((----(........))))))..(((((.(((((((((.(((...(((..(((.....))).)))...))).)))))))))))))).)))....)))......- ( -23.70, z-score = -1.26, R) >droEre2.scaffold_4690 6245189 103 + 18748788 AUGUUAAUGCC-AUUGGU----UUCCACAGUGACCGAAAUUGAUGUCAAUUGAUAUUGAUAAUCCAUGCAUUAUGCGCGAUAAACAAAGUUAAUUAAAUCAAAGGCUC------------ ........(((-.(((((----(........))))))..(((((...(((((((.(((...(((..(((.....))).)))...))).)))))))..))))).)))..------------ ( -21.70, z-score = -1.09, R) >droPer1.super_17 168958 107 + 1930428 CUGCUAAGAUUCCUUGGUCCACCCCCCCCGUUACCGAAAUUGACGUCAAUUGAUAUUGAUAAUCCAUGCAUUAUGCGCGAUAAACAAAGUUAAUUGAAUCAAAGGCG------------- ..(((..(((((.(((((..((.......)).)))))(((((((((((((....)))))).(((..(((.....))).))).......))))))))))))...))).------------- ( -20.10, z-score = -1.27, R) >dp4.chrXL_group1e 7717325 106 + 12523060 CUGCUAAGAUUCCUUGGUCCA-CCCCCCCGCUACCGAAAUUGAUGUCAAUUGAUAUUGAUAAUCCAUGCAUUAUGCGCGAUAAACAAAGUUAAUUGAAUCAAAGGCG------------- ..((((((....))))))...-......((((.......(((((.(((((((((.(((...(((..(((.....))).)))...))).)))))))))))))).))))------------- ( -20.60, z-score = -1.34, R) >consensus AUGUUAAUGCU_AUUGGU____UGCCACAGUGACCAAAAUUGAUGUCAAUUGAUAUUGAUAAUCCAUGCAUUAUGCGCGAUAAACAAAGUUAAUUGAAUCAAAGGCGC__A_CC_ACAA_ ..(((........(((((..............)))))..(((((.(((((((((.(((...(((..(((.....))).)))...))).)))))))))))))).))).............. (-17.86 = -17.68 + -0.19)

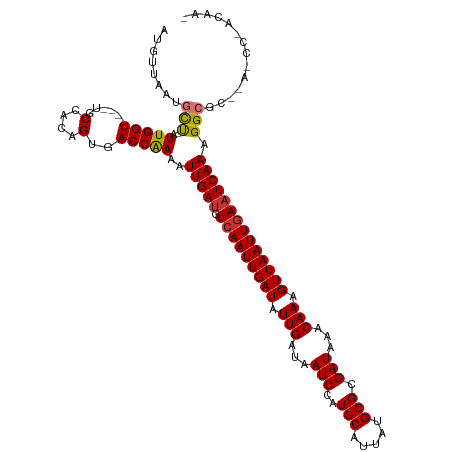

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:43:49 2011