
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 11,398,033 – 11,398,135 |
| Length | 102 |
| Max. P | 0.529884 |

| Location | 11,398,033 – 11,398,135 |
|---|---|
| Length | 102 |
| Sequences | 9 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 72.40 |
| Shannon entropy | 0.55847 |
| G+C content | 0.49315 |
| Mean single sequence MFE | -36.18 |
| Consensus MFE | -9.83 |
| Energy contribution | -8.73 |
| Covariance contribution | -1.10 |
| Combinations/Pair | 1.71 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.27 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.529884 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 11398033 102 - 23011544 GCUUGGUCCAUUUGCAUAUG-----UCGUCG-AUGCGAUCGGCAGGGAGUUGUCCAUUUUAUAAUUGGAUAGUUUGUCACCGCGUUGGAGUAUCUGGGAGUGGGGCCA----- ...((((((((((.((.(((-----.(..((-(((((...((((((...(((((((.........)))))))))))))..)))))))..)))).)).)))).))))))----- ( -35.00, z-score = -1.77, R) >droSim1.chr2L 11206522 102 - 22036055 GCUUGCCCCAUUUGCAUAUG-----UCGUCG-AUGCGAUCGGCAAGGAGUUGUCCAUUUUAUAAUUGGAUAGUUUGUCACCGCGUUGGAGUAUCUGCAAGUGGGGCCA----- ....((((((((((((.(((-----.(..((-(((((...((((((...(((((((.........)))))))))))))..)))))))..)))).))))))))))))..----- ( -45.00, z-score = -5.05, R) >droSec1.super_3 6792339 102 - 7220098 GCUUGCCCCAUUUGCAUAUG-----UCGUCG-AUGCGAUCGGCAGGGAGUUGUCCAUUUUAUAAUAGGAUAGUUUGUCACCGCGUUGGAGUAUCUGCAAGUGACGCCA----- ....((..((((((((.(((-----.(..((-(((((...((((((...(((((((((....))).))))))))))))..)))))))..)))).))))))))..))..----- ( -31.50, z-score = -1.15, R) >droYak2.chr2L 7795997 102 - 22324452 GCUUGCCCCAUUUGCAUAUG-----UCGUCG-AUGCGAUCGGCAGGGAGUUGUCCAUUUUAUAAUUGGAUAGUUUGUCACCGCGUUGGAGUAUCUGCAAGUGGGGCCA----- ....((((((((((((.(((-----.(..((-(((((...((((((...(((((((.........)))))))))))))..)))))))..)))).))))))))))))..----- ( -44.60, z-score = -4.59, R) >droEre2.scaffold_4929 12599490 102 + 26641161 GCUUGCCCCAUUUGCAUAUG-----UCGUCG-AUGCGAUCGGCAGGGAGUUGUCCAUUUUAUAAUUGGAUAGUUUGUCACCGCGCCGGAGUAGCUGCAAGUGGGGCCA----- ....((((((((((((...(-----((((..-..)))))((((.((((.(((((((.........)))))))....)).))..)))).......))))))))))))..----- ( -41.10, z-score = -2.76, R) >droAna3.scaffold_12916 12106846 101 + 16180835 GCUUG-CCCAUUUGCAUAUG-----UCGUCG-AUGCGAUCAGUUGGGAGUUGUCCAUUUUAUAAUUGGAUAGUUUGUCACCACGCCGGGGUAUCCAAAGUUGUGGCCA----- .....-((((.((((((.((-----....))-)))))).....))))..(((((((.........)))))))...(((((.((....((....))...)).)))))..----- ( -24.50, z-score = 0.78, R) >droPer1.super_1 8296508 90 - 10282868 GCUUGCCCCAUUUGCAUACG-----UCUGCGUUUGGGGCCUGCUCGGAAUUGUCCAUUUUAUAAUUGGAUAGUUUGUCAUUG---GGAGGCAGGAGCA--------------- ((((((((((..((((....-----..))))..))))))(((((..((((((((((.........)))))))))).((....---.))))))))))).--------------- ( -36.90, z-score = -3.40, R) >dp4.chr4_group3 6848610 113 - 11692001 GCUUGCCCCAUUUGCAUACGUCUGCUCUGCGUUUGGGGCCUGCUCGGAAUUGUCCAUUUUAUAAUUGGAUAGUUUGUCACCGGCUGGGGACUGGGGUCUGGGAGGCAGGAGCA ((((((((((..((((...........))))..))))))(((((..((((((((((.........)))))))))).((.(((((((....)...)).))))))))))))))). ( -44.00, z-score = -1.38, R) >droGri2.scaffold_15126 7216368 89 + 8399593 ----GCUCCUGUAUUUUAC------CUGAGGCCAAGAAUUGAGUAAGAGCUGUUGACUUUAUAAUUGGAUAGUUUGUUACUGAGCAGUGGUCGGGGUAA-------------- ----...........((((------((..(((((....((.(((((((((((((.(.........).)))))))).))))).))...))))).))))))-------------- ( -23.00, z-score = -0.94, R) >consensus GCUUGCCCCAUUUGCAUAUG_____UCGUCG_AUGCGAUCGGCAGGGAGUUGUCCAUUUUAUAAUUGGAUAGUUUGUCACCGCGUUGGAGUAUCUGCAAGUGGGGCCA_____ ...........(((((.................)))))...((((.((((((((((.........))))))))))....((.....)).....))))................ ( -9.83 = -8.73 + -1.10)


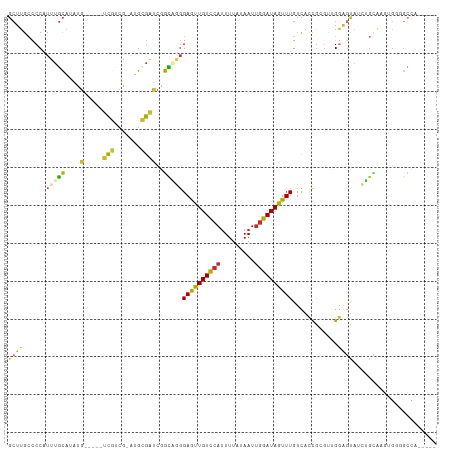
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:32:23 2011