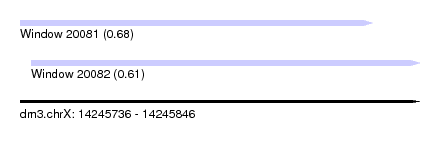

| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,245,736 – 14,245,846 |
| Length | 110 |
| Max. P | 0.683165 |

| Location | 14,245,736 – 14,245,833 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 71.51 |
| Shannon entropy | 0.49140 |
| G+C content | 0.59135 |
| Mean single sequence MFE | -40.76 |
| Consensus MFE | -19.50 |
| Energy contribution | -21.14 |
| Covariance contribution | 1.64 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.683165 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
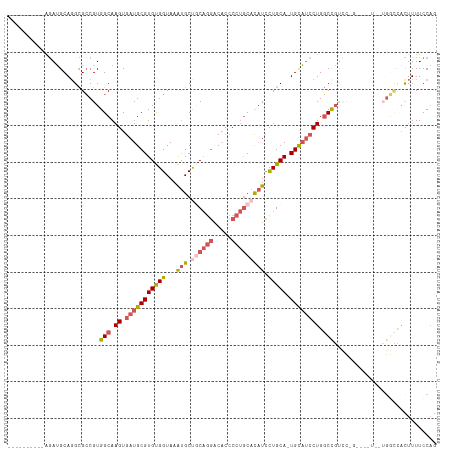
>dm3.chrX 14245736 97 + 22422827 ----------AGAUGCAGGCGCCGUGGCAAGUGAUGCGUGUGGUAAAUGCCGCAGGACACCCCUGCACAUCCUGCA-UGCAUCCUGGCCGUCC---------UGGCCACUUUUCCAG ----------.((...((..((((((((.((.((((((((((((....)))(((((.....))))).......)))-)))))))).))))...---------))))..))..))... ( -37.70, z-score = -1.04, R) >droSec1.super_20 848426 106 + 1148123 ----------AGAUGCAUGCGCCGUGGCAAGUGAUGCGUGUGGUAAAUGCUGCAGGACACCCCUGCACAUCCUGCA-UGCAUCCUGGCCGUCCCGGCCAUCCUGGCCACUUUUCCAG ----------....((....)).(((((.((.((((((((..(...(((.((((((.....))))))))).)..))-))))))))(((((...)))))......)))))........ ( -45.60, z-score = -2.69, R) >droYak2.chrX 8488401 106 + 21770863 ----------AGAUGCAGGCGCCGUGGCAAGUGAUGCGUGUGGUAAAUGCUGCAGGACACCCCUGCACAUCCUGCA-UGCAUCCUGGCCGUCCUGCUCGUCCUGGCCACUUUUCCAG ----------.((.(((((.((...(((.((.((((((((..(...(((.((((((.....))))))))).)..))-)))))))).))))))))))))...((((........)))) ( -45.80, z-score = -2.57, R) >droEre2.scaffold_4690 6051609 88 - 18748788 ----------AGAUGCAGGCGCCGUGGCAAGUGAUGCGUGUGGUAAAUGCUCCAGGACACCCCUGCACAUCCUGCA-UGCAUCCUGGCC------------------ACUUUUCCAG ----------.......((....(((((.((.((((((((..(...(((...((((.....))))..))).)..))-)))))))).)))------------------))....)).. ( -33.60, z-score = -1.55, R) >droAna3.scaffold_13248 1939410 114 + 4840945 GGCUGAAGGGGCAGGCAGGUGGCGUGGCAAGUGAUGCGUGCGGUAAGCGCCACCAUCCUGGCC-ACUUUUGUUGCAAUGUCCGCUGUCUGUGAUGGCGUUUUGGUUUUUUUUUCC-- (((..(((.(.((.(((((..(((.((((((.((((.(((((.....)).))))))))).((.-((....)).))..)))))))..)))))..)).).)))..))).........-- ( -41.10, z-score = -0.13, R) >consensus __________AGAUGCAGGCGCCGUGGCAAGUGAUGCGUGUGGUAAAUGCUGCAGGACACCCCUGCACAUCCUGCA_UGCAUCCUGGCCGUCC_G____U__UGGCCACUUUUCCAG .........................(((.((.(((((((((((...(((.((((((.....))))))))).))))).)))))))).)))............................ (-19.50 = -21.14 + 1.64)
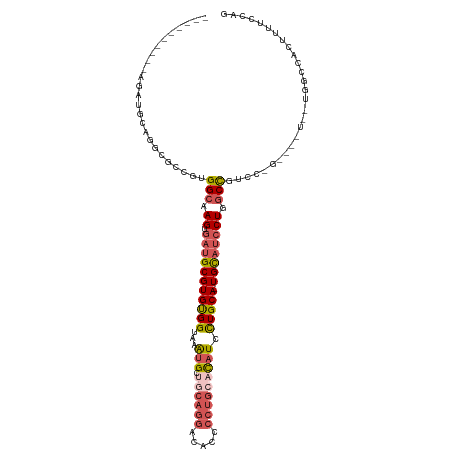


| Location | 14,245,739 – 14,245,846 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 88.50 |
| Shannon entropy | 0.18575 |
| G+C content | 0.57845 |
| Mean single sequence MFE | -42.88 |
| Consensus MFE | -32.72 |
| Energy contribution | -34.98 |
| Covariance contribution | 2.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.606348 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14245739 107 + 22422827 UGCAGGCGCCGUGGCAAGUGAUGCGUGUGGUAAAUGCCGCAGGACACCCCUGCACAUCCUGCAUGCAUCCUGGCCGUC---------CUGGCCACUUUUCCAGCACUGUUCGUUUA .((((((((....))((((((((((((((((....)))(((((.....))))).......)))))))))..((((...---------..)))))))).....)).))))....... ( -41.80, z-score = -1.32, R) >droSec1.super_20 848429 116 + 1148123 UGCAUGCGCCGUGGCAAGUGAUGCGUGUGGUAAAUGCUGCAGGACACCCCUGCACAUCCUGCAUGCAUCCUGGCCGUCCCGGCCAUCCUGGCCACUUUUCCAGAACUGUUCGUUUA .((....)).(((((.((.((((((((..(...(((.((((((.....))))))))).)..))))))))))(((((...)))))......))))).......((((.....)))). ( -46.30, z-score = -2.47, R) >droYak2.chrX 8488404 116 + 21770863 UGCAGGCGCCGUGGCAAGUGAUGCGUGUGGUAAAUGCUGCAGGACACCCCUGCACAUCCUGCAUGCAUCCUGGCCGUCCUGCUCGUCCUGGCCACUUUUCCAGCACUGUUCGUUUA .(((((.((...(((.((.((((((((..(...(((.((((((.....))))))))).)..)))))))))).))))))))))..((.((((........)))).)).......... ( -46.70, z-score = -2.04, R) >droEre2.scaffold_4690 6051612 98 - 18748788 UGCAGGCGCCGUGGCAAGUGAUGCGUGUGGUAAAUGCUCCAGGACACCCCUGCACAUCCUGCAUGCAUC------------------CUGGCCACUUUUCCAGCACUGUUCGUUUA .((((((...(((((.((.((((((((..(...(((...((((.....))))..))).)..))))))))------------------)).))))).......)).))))....... ( -36.70, z-score = -1.63, R) >consensus UGCAGGCGCCGUGGCAAGUGAUGCGUGUGGUAAAUGCUGCAGGACACCCCUGCACAUCCUGCAUGCAUCCUGGCCGUC_________CUGGCCACUUUUCCAGCACUGUUCGUUUA .((((((((....))((((((((((((..(...(((.((((((.....))))))))).)..))))))))..(((((............))))))))).....)).))))....... (-32.72 = -34.98 + 2.25)
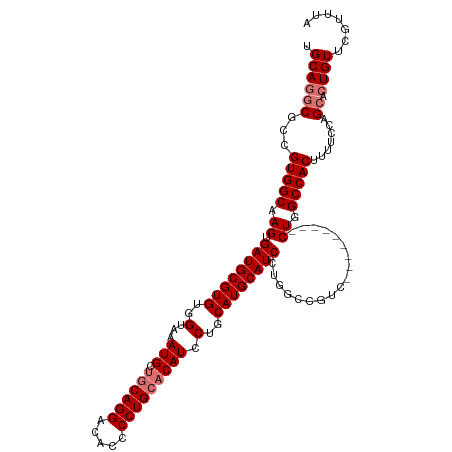



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:42:55 2011