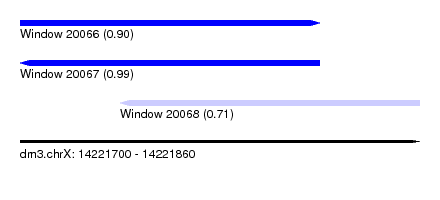
| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,221,700 – 14,221,860 |
| Length | 160 |
| Max. P | 0.985668 |

| Location | 14,221,700 – 14,221,820 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.92 |
| Shannon entropy | 0.11127 |
| G+C content | 0.39667 |
| Mean single sequence MFE | -34.16 |
| Consensus MFE | -28.24 |
| Energy contribution | -28.20 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.83 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.15 |
| SVM RNA-class probability | 0.900210 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14221700 120 + 22422827 AGCCAAUAUAUCUUUCACAAGAUAUAAGUGCAUGGAUGUGUAUUUUCCGGCCGUAUAGCCAUCAAAAAGCGGAUGCUUCUUGUUGGAAAUCCGGCAAUUAGAGGAAAUUCGUUUGUGUAU ......((((((((....)))))))).(((((..((((...(((((((.((((.((..(((.(((.((((....)))).))).)))..)).)))).....).)))))).))))..))))) ( -36.10, z-score = -2.97, R) >droEre2.scaffold_4690 6028512 120 - 18748788 AGGGAAUAUAUCUUUCACAAGACACAAGUGCUUGGAUGUAUACUUUCCGGCCGUAGAGCCAUCAAAAAGCGGAUGCUUCUUGUUGGAAAUCCGGCAAUUAGAGGAAAUUCGUUUGUGUAU .((((((((((...((.((((.((....)))))))).))))).)))))(((......)))......((((((((.(((((((((((....))))))...)))))..))))))))...... ( -34.40, z-score = -1.92, R) >droYak2.chrX 8464974 120 + 21770863 AGAAAAUAUAUCUGUCACACGAUAAAAGUGCUUGGAUGUAUACUUUCUGGCCGUAGAGCCAUCAAAAAGCGGAUGCUUCUUGUUGGAAUGCCGGCAAUUAGAGGAAAUUCGUUUGUGUAU ((((((((((((((...(((.......)))..)))))))))..)))))(((......)))......((((((((.(((((((((((....))))))...)))))..))))))))...... ( -31.90, z-score = -1.52, R) >droSec1.super_20 824520 120 + 1148123 AGUCAAUAUAUCUUUCACAAGAUAUAAGUGCAUGGAUGUGUACUUUCCGGCCGUAUAGACAUCAAAAAGCGGAUGCUUCUUGUUGGAAAUCCGGCAAUUAGAGGAAAUUCGUUUGUGUAU .(((..((((((((....))))))))(((((((....))))))).............)))......((((((((.(((((((((((....))))))...)))))..))))))))...... ( -33.00, z-score = -2.32, R) >droSim1.chrX 10916149 120 + 17042790 AGUCAAUAUAUCUUUCACAAGAUAUAAGUGCAUGGAUGUGUACUUUCCGGCCGUAUAGCCAUCAAAAAGCGGAUGCUUCUUGUUGGAAAUCCGGCAAUUAGAGGAAAUUCGUUUGUGUAU ......((((((((....)))))))).(((((..((((.....(((((.((((.((..(((.(((.((((....)))).))).)))..)).)))).......)))))..))))..))))) ( -35.40, z-score = -2.94, R) >consensus AGUCAAUAUAUCUUUCACAAGAUAUAAGUGCAUGGAUGUGUACUUUCCGGCCGUAUAGCCAUCAAAAAGCGGAUGCUUCUUGUUGGAAAUCCGGCAAUUAGAGGAAAUUCGUUUGUGUAU .....((((((((..(((.........)))...)))))))).......(((......)))......((((((((.(((((((((((....))))))...)))))..))))))))...... (-28.24 = -28.20 + -0.04)


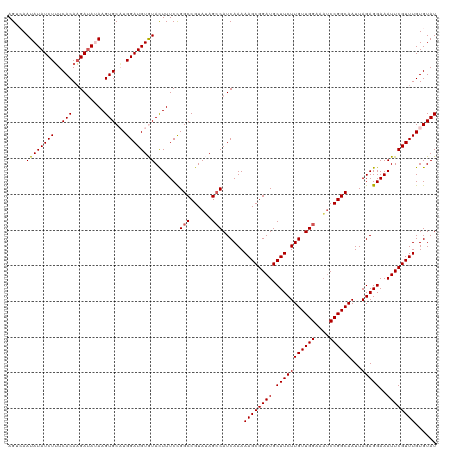
| Location | 14,221,700 – 14,221,820 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.92 |
| Shannon entropy | 0.11127 |
| G+C content | 0.39667 |
| Mean single sequence MFE | -30.03 |
| Consensus MFE | -24.54 |
| Energy contribution | -25.26 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.07 |
| Mean z-score | -3.12 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.21 |
| SVM RNA-class probability | 0.985668 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14221700 120 - 22422827 AUACACAAACGAAUUUCCUCUAAUUGCCGGAUUUCCAACAAGAAGCAUCCGCUUUUUGAUGGCUAUACGGCCGGAAAAUACACAUCCAUGCACUUAUAUCUUGUGAAAGAUAUAUUGGCU .........................(((((((((((..((((((((....))))))))..((((....)))))))).......)))).......((((((((....))))))))..))). ( -32.51, z-score = -3.83, R) >droEre2.scaffold_4690 6028512 120 + 18748788 AUACACAAACGAAUUUCCUCUAAUUGCCGGAUUUCCAACAAGAAGCAUCCGCUUUUUGAUGGCUCUACGGCCGGAAAGUAUACAUCCAAGCACUUGUGUCUUGUGAAAGAUAUAUUCCCU .........................((((.....(((.((((((((....)))))))).))).....)))).((((.((((...((((((((....)).)))).))...)))).)))).. ( -32.10, z-score = -3.07, R) >droYak2.chrX 8464974 120 - 21770863 AUACACAAACGAAUUUCCUCUAAUUGCCGGCAUUCCAACAAGAAGCAUCCGCUUUUUGAUGGCUCUACGGCCAGAAAGUAUACAUCCAAGCACUUUUAUCGUGUGACAGAUAUAUUUUCU .........................((((.....(((.((((((((....)))))))).))).....)))).((((((((((.......((((.......))))......)))))))))) ( -30.52, z-score = -3.55, R) >droSec1.super_20 824520 120 - 1148123 AUACACAAACGAAUUUCCUCUAAUUGCCGGAUUUCCAACAAGAAGCAUCCGCUUUUUGAUGUCUAUACGGCCGGAAAGUACACAUCCAUGCACUUAUAUCUUGUGAAAGAUAUAUUGACU .....(((.....(((((.......((((......((.((((((((....)))))))).))......)))).))))).................((((((((....)))))))))))... ( -25.40, z-score = -2.06, R) >droSim1.chrX 10916149 120 - 17042790 AUACACAAACGAAUUUCCUCUAAUUGCCGGAUUUCCAACAAGAAGCAUCCGCUUUUUGAUGGCUAUACGGCCGGAAAGUACACAUCCAUGCACUUAUAUCUUGUGAAAGAUAUAUUGACU .....(((................(((.((((((((..((((((((....))))))))..((((....)))))))).......))))..)))..((((((((....)))))))))))... ( -29.61, z-score = -3.11, R) >consensus AUACACAAACGAAUUUCCUCUAAUUGCCGGAUUUCCAACAAGAAGCAUCCGCUUUUUGAUGGCUAUACGGCCGGAAAGUACACAUCCAUGCACUUAUAUCUUGUGAAAGAUAUAUUGACU .............(((((.......((((.....(((.((((((((....)))))))).))).....)))).))))).................((((((((....))))))))...... (-24.54 = -25.26 + 0.72)


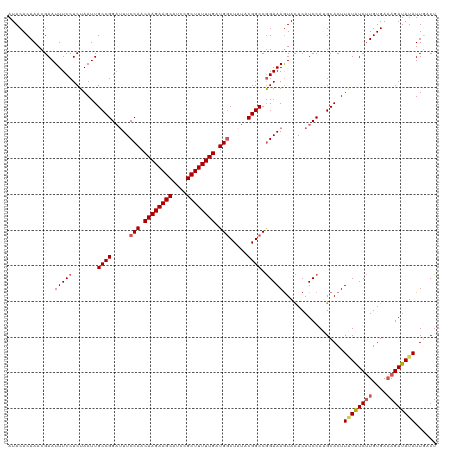
| Location | 14,221,740 – 14,221,860 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.17 |
| Shannon entropy | 0.05020 |
| G+C content | 0.40500 |
| Mean single sequence MFE | -22.72 |
| Consensus MFE | -22.26 |
| Energy contribution | -22.46 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.15 |
| Structure conservation index | 0.98 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.712272 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14221740 120 - 22422827 CAGAGCGAUCAAAAGUAAACAAAAGGAACCACACCAAUUAAUACACAAACGAAUUUCCUCUAAUUGCCGGAUUUCCAACAAGAAGCAUCCGCUUUUUGAUGGCUAUACGGCCGGAAAAUA .((((.((......(....)....((.......))....................))))))....((((.((..(((.((((((((....)))))))).)))..)).))))......... ( -23.60, z-score = -1.68, R) >droEre2.scaffold_4690 6028552 120 + 18748788 CAGAGCGAUAAAAAGUAAACAAAAGGAACCAUACCAAUUAAUACACAAACGAAUUUCCUCUAAUUGCCGGAUUUCCAACAAGAAGCAUCCGCUUUUUGAUGGCUCUACGGCCGGAAAGUA .((((.((......(....)....((.......))....................))))))....((((.....(((.((((((((....)))))))).))).....))))......... ( -23.30, z-score = -1.07, R) >droYak2.chrX 8465014 120 - 21770863 CAGAGCGAUCAAAAGUAAACAAAAGGAACCACACCAAUUAAUACACAAACGAAUUUCCUCUAAUUGCCGGCAUUCCAACAAGAAGCAUCCGCUUUUUGAUGGCUCUACGGCCAGAAAGUA .((((.((......(....)....((.......))....................))))))....((((.....(((.((((((((....)))))))).))).....))))......... ( -23.30, z-score = -1.39, R) >droSec1.super_20 824560 120 - 1148123 CAGAGCGAUCAAAAGUAAACAAAAGGAACCACACCAAUUAAUACACAAACGAAUUUCCUCUAAUUGCCGGAUUUCCAACAAGAAGCAUCCGCUUUUUGAUGUCUAUACGGCCGGAAAGUA .((((.((......(....)....((.......))....................))))))....((((......((.((((((((....)))))))).))......))))......... ( -19.80, z-score = -0.33, R) >droSim1.chrX 10916189 120 - 17042790 CAGAGCGAUCAAAAGUAAACAAAAGGAACCACACCAAUUAAUACACAAACGAAUUUCCUCUAAUUGCCGGAUUUCCAACAAGAAGCAUCCGCUUUUUGAUGGCUAUACGGCCGGAAAGUA .((((.((......(....)....((.......))....................))))))....((((.((..(((.((((((((....)))))))).)))..)).))))......... ( -23.60, z-score = -1.31, R) >consensus CAGAGCGAUCAAAAGUAAACAAAAGGAACCACACCAAUUAAUACACAAACGAAUUUCCUCUAAUUGCCGGAUUUCCAACAAGAAGCAUCCGCUUUUUGAUGGCUAUACGGCCGGAAAGUA .((((.((......(....)....((.......))....................))))))....((((.....(((.((((((((....)))))))).))).....))))......... (-22.26 = -22.46 + 0.20)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:42:44 2011