
| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,116,714 – 14,116,810 |
| Length | 96 |
| Max. P | 0.659278 |

| Location | 14,116,714 – 14,116,810 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 69.87 |
| Shannon entropy | 0.57708 |
| G+C content | 0.56689 |
| Mean single sequence MFE | -31.74 |
| Consensus MFE | -14.43 |
| Energy contribution | -14.64 |
| Covariance contribution | 0.21 |
| Combinations/Pair | 1.52 |
| Mean z-score | -1.29 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.659278 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14116714 96 - 22422827 ----UAACAUAAC-GCGGCUAACGGCUACAAAUGUGACCCACAUCCCGUCCAGCUUUGUGCUGCAGUCGUUGACC-----CUGCAGUGGGUCAAGGCGCCUCUCUG ----.........-(((.(((((((((....(((((...)))))......((((.....)))).)))))))((((-----(......))))).)).)))....... ( -30.80, z-score = -1.19, R) >droGri2.scaffold_15081 2998795 85 + 4274704 --------GAGAAGGACGCCGUCGGUCAUGAAUUUAAUGCGCGUCGC---UGGCGUCACAUUGCCCUCGAAG---------CGUAUCAAAU-AGGACACCUCGCUG --------(((...(((...))).(((....((((.((((((.(((.---.((((......))))..))).)---------))))).))))-..)))..))).... ( -25.20, z-score = -0.30, R) >droSim1.chrU 2037587 96 + 15797150 ----UAACAUAAC-GCGACUAGCGGCUACAAAUGUGACCCACAUCCCAUCCAGCUUUGCGCUGCAGUCGUUGACC-----CUGCAGUGGGUCAAGGCGCCUCUCUG ----........(-((((((.(((((.....(((((...)))))........((...))))))))))).((((((-----(......))))))).)))........ ( -30.60, z-score = -1.52, R) >droSec1.super_20 720581 96 - 1148123 ----UAGCAUAAC-GCGACUAGCGGCUACAAAUGUGACCCACAUCCCAUCCAGCUUUGCGCUGCAGUCGUUGACC-----CUGCAGUGGGUCAAGGCGCCUCUCUG ----..((.....-((((((.(((((.....(((((...)))))........((...)))))))))))))(((((-----(......))))))..))......... ( -31.10, z-score = -1.06, R) >droYak2.chrX 8369969 96 - 21770863 ----UCACAUAAC-ACGGCUAACGGUCACAAAUGUGACCCACAUCUCGUCCAGCUUUGCGCUGCAGUCGUUGACC-----CUGCAGUGGGUCAAGGCGUUUCUCCG ----.........-.(((..(((((((((....))))))...........((((.....))))..(((.((((((-----(......)))))))))))))...))) ( -32.60, z-score = -2.39, R) >droEre2.scaffold_4690 5935079 96 + 18748788 ----UCGCAUAAC-CCGACUAACGUUUACAAAUGUGACCCACAUCCCGCCCAGCUUUGCGCUGCAGUCGUUGACC-----CUGCAGUGGGUCAAGGCGUUUCUCUG ----((((((...-..(((....))).....)))))).........((((((((.....))))......((((((-----(......)))))))))))........ ( -28.70, z-score = -1.87, R) >droAna3.scaffold_12903 328264 106 + 802071 GGGGUGGCGCAGUGGGGGUCAAGUGGCGGGUAUGUGACCCACAUCCCGUCCAGCUUUUUCCUGCAGUCGUUGACCGCUGUGUGCAGUGGGUCAAGGCAUUUGCUGG .....(((...(..((((..((((((((((.(((((...))))))))).))).))).))))..).(((.((((((((((....)))).)))))))))....))).. ( -43.20, z-score = -0.69, R) >consensus ____UAACAUAAC_GCGGCUAACGGCUACAAAUGUGACCCACAUCCCGUCCAGCUUUGCGCUGCAGUCGUUGACC_____CUGCAGUGGGUCAAGGCGCCUCUCUG ..............(((.((...(....).....((((((((.........(((.....)))((((..............)))).)))))))))).)))....... (-14.43 = -14.64 + 0.21)


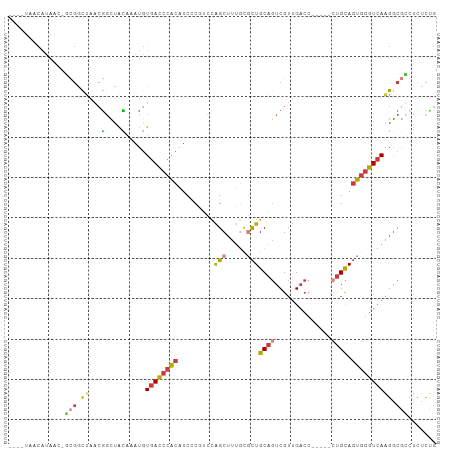
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:42:28 2011