
| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,113,326 – 14,113,420 |
| Length | 94 |
| Max. P | 0.768505 |

| Location | 14,113,326 – 14,113,420 |
|---|---|
| Length | 94 |
| Sequences | 8 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 72.36 |
| Shannon entropy | 0.50592 |
| G+C content | 0.35441 |
| Mean single sequence MFE | -22.62 |
| Consensus MFE | -12.54 |
| Energy contribution | -11.76 |
| Covariance contribution | -0.78 |
| Combinations/Pair | 1.47 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.768505 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
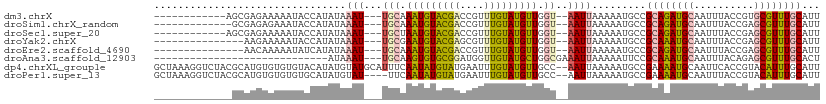
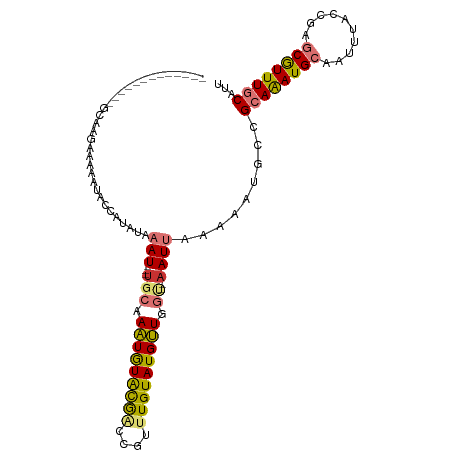
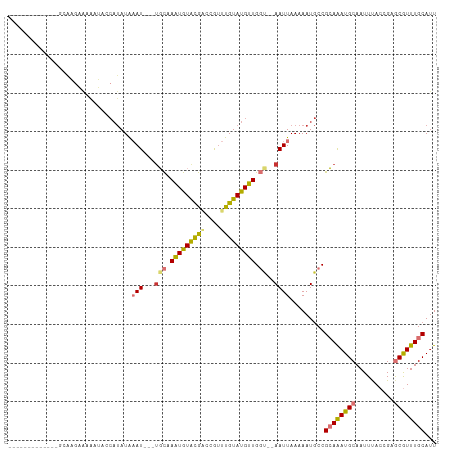
>dm3.chrX 14113326 94 - 22422827 ------------AGCGAGAAAAAUACCAUAUAAAU---UGCAAAUGUACGACCGUUUGUAUGUUGGU--AAUUAAAAAUGCCGCAGAUGCAAUUUACCGUGCGUUUGCAUU ------------.......................---((((((((((((.....((((((.(((((--(........))))).).)))))).....)))))))))))).. ( -26.50, z-score = -2.82, R) >droSim1.chrX_random 3784550 93 - 5698898 -------------GCGAGAGAAAUACCAUAUAAAU---UGCAAAUGUACGACCGUUUGUAUGUUGGU--AAUUAAAAAUGCCGCAGAUGCAAUUUACCGAGCGUUUGCAUU -------------(((....((.(((((.......---((((((((......))))))))...))))--).)).....))).((((((((..........))))))))... ( -22.40, z-score = -1.73, R) >droSec1.super_20 717237 94 - 1148123 ------------AGCGAGAAAAAUACCAUAUAAAU---UGCUAAUGUACGACCGUUUGUAUGUUGGU--AAUUAAAAAUGCCGCAGAUGCAAUUUACCGAGCGUUUGCAUU ------------....................(((---(((((((((((((....))))))))))))--)))).........((((((((..........))))))))... ( -24.80, z-score = -2.68, R) >droYak2.chrX 8366702 91 - 21770863 ---------------AAGAAAAAUACCAUAUAAAU---UGCGAAUGUACGAGCGUUUGUAUGUUGGU--AAUUAAAAAUGCCGCAAAUGCAAUUUACCGAGCGUUUGCAUU ---------------........(((((.......---(((((((((....)))))))))...))))--)............((((((((..........))))))))... ( -22.80, z-score = -2.01, R) >droEre2.scaffold_4690 5931509 91 + 18748788 ---------------AACAAAAAUAUCAUAUAAAU---UGCAAAUGUACGACCGUUUGUAUGUUGGU--AAUUAAAAAUGCCGCAGAUGCAAUUUACCGAGCGUUUGCAUU ---------------....................---(((((((((.((.....((((((.(((((--(........))))).).)))))).....)).))))))))).. ( -22.20, z-score = -2.00, R) >droAna3.scaffold_12903 325625 79 + 802071 -----------------------------AUAAAU---UGCAAGUGUGCGGAUGGUUGUAUGCUGGCGAAAUUAAAAAUUCCGCAAAUGCAAUUUACAGAGCGUUUGCACU -----------------------------.....(---(((.(((((((((....))))))))).)))).............((((((((..........))))))))... ( -21.90, z-score = -1.36, R) >dp4.chrXL_group1e 8458912 109 - 12523060 GCUAAAGGUCUACGCAUGUGUGUGUACAUAUGUAUGCAUUUCAAUAUGUAUGAAUUUGUAUGUUGCC--AAUUAAAAAUGCCGAAAAUGCAAUUCACCGUACAUUUGCAUU ((...........(((((..(((....)))..)))))........(((((((...((((((.(((.(--(........)).)))..)))))).....)))))))..))... ( -20.90, z-score = 0.97, R) >droPer1.super_13 2177806 105 + 2293547 GCUAAAGGUCUACGCAUGUGUGUGUGCAUAUGUAU----UUCAAUAUGUAUGAAUUUGUAUGUUGCC--AAUUAAAAAUGCCGAAAAUGCAAUUUACCGUACAUUUGCAUU ((...........((((((((....))))))))..----......(((((((...((((((.(((.(--(........)).)))..)))))).....)))))))..))... ( -19.50, z-score = 0.97, R) >consensus _____________GCAAGAAAAAUACCAUAUAAAU___UGCAAAUGUACGACCGUUUGUAUGUUGGU__AAUUAAAAAUGCCGCAAAUGCAAUUUACCGAGCGUUUGCAUU ..........................................(((((((((....)))))))))..................((((((((..........))))))))... (-12.54 = -11.76 + -0.78)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:42:23 2011