
| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,109,344 – 14,109,497 |
| Length | 153 |
| Max. P | 0.821804 |

| Location | 14,109,344 – 14,109,497 |
|---|---|
| Length | 153 |
| Sequences | 5 |
| Columns | 174 |
| Reading direction | forward |
| Mean pairwise identity | 69.12 |
| Shannon entropy | 0.50114 |
| G+C content | 0.39582 |
| Mean single sequence MFE | -27.11 |
| Consensus MFE | -16.53 |
| Energy contribution | -17.09 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.821804 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 14109344 153 + 22422827 AAUCAUAUAAUUUACUUAAGUAUAUUCCUUAAAA-UGUCGAAUACUCGCUAGGUGCUACCUUUAUGACCGCUGCUGUACUCCCCCCCGUUCAACCACCUGCAGUUUCUCUAAUUGCAGUUUUUGCAAUUUGCUUUUAUUUAUGAACAUAUGUAC-------------------- ...(((((..........((((((..........-....((....))((..(((.(.........))))...)))))))).......(((((.....((((((((.....)))))))).....((.....)).........))))))))))...-------------------- ( -24.20, z-score = -1.18, R) >droSec1.super_20 712877 140 + 1148123 -----------------AGCCGUACAGCUUAGUUCCAAAUAAU-CUCGCUGGGUGCUACCUUUUUGACCGCUGCCGUACUCCCCCCCGUUCAACCACCUGCAGUUUUUCUAAUUGCAGUUUUUACAAUUUGCUUUUAUUUAUGAACAUGCAUAUGUAC---------------- -----------------....(((((((...((((.((((((.-...((.((((((...................))))))................((((((((.....))))))))............))..))))))..))))..))...)))))---------------- ( -23.81, z-score = -0.30, R) >droSim1.chrX_random 3782084 140 + 5698898 -----------------AAGCGUACAGCUUAGUUCCAAAUAAU-CUCGCUGGGUGCUACCUUUUUGACCGCUGCUGUACUCCCCCCCGUUCAACCACCUGCAGUUUUUCUAAUUGCAGUUUUUGCAAUUUGCUUUUAUUUAUGAACAUGCAUAUGUAC---------------- -----------------..(((((((((..(((..((((....-...((.....))......))))...))))))))))........(((((.....((((((((.....)))))))).....((.....)).........)))))..))........---------------- ( -31.02, z-score = -1.97, R) >droYak2.chrX 8362893 115 + 21770863 -----------------------------------UGUUGAUUAUUUGUUGAGCGCUUCCAUUUUGACCGCUGCUGUACUCCUCCCCAUUCAACCACCUGCAGUUUCUCUAAUUGCAGU----GCAAUUUGCUUUUAUUUAUGAACAUAUGUAC-------------------- -----------------------------------.(((((......(..(((..(...................)..)))..).....)))))...((((((((.....)))))))).----((.....))......................-------------------- ( -15.81, z-score = 0.34, R) >droEre2.scaffold_4690 5927542 168 - 18748788 --GUUUCUAAUGUAUUUUUAUACAUCAUUUAAAAGUGUUGAGUAUUUGUUGAGUGCUACCAUUCUGACCGCUGCUGUACUCCUCCCCGUUCAACCACCUGCAGUUUCUCUAAUUGCAGU----GCAAUUUGCUUUUAUUUAUGAACAUAUGUAUGUACAUAUGUGCAUGUGUAC --........(((((....)))))((((.(((((((((((((.....(..((((((...................))))))..)....))))))(((.(((((((.....)))))))))----)......)))))))...))))((((((((((((....)))))))))))).. ( -40.71, z-score = -2.98, R) >consensus _________________AAGCAUACAGCUUAAAA_UGUAGAAUACUCGCUGGGUGCUACCUUUUUGACCGCUGCUGUACUCCCCCCCGUUCAACCACCUGCAGUUUCUCUAAUUGCAGUUUUUGCAAUUUGCUUUUAUUUAUGAACAUAUGUAUGUAC________________ ..................................................((((((...................))))))......(((((.....((((((((.....)))))))).....((.....)).........)))))............................ (-16.53 = -17.09 + 0.56)

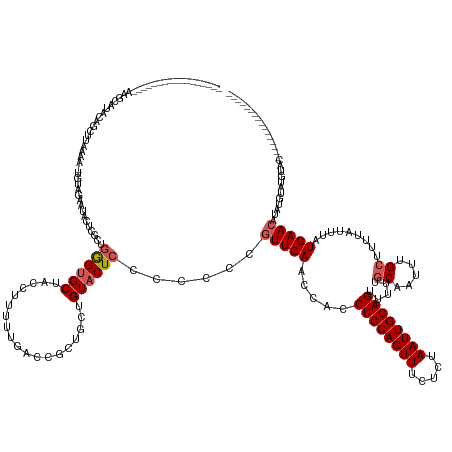

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:42:22 2011