
| Sequence ID | dm3.chrX |
|---|---|
| Location | 14,081,917 – 14,082,013 |
| Length | 96 |
| Max. P | 0.692765 |

| Location | 14,081,917 – 14,082,013 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 61.05 |
| Shannon entropy | 0.79419 |
| G+C content | 0.44383 |
| Mean single sequence MFE | -25.50 |
| Consensus MFE | -9.14 |
| Energy contribution | -8.59 |
| Covariance contribution | -0.56 |
| Combinations/Pair | 1.96 |
| Mean z-score | -1.00 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.692765 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
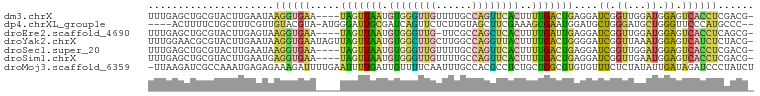
>dm3.chrX 14081917 96 - 22422827 UUUGAGCUGCGUACUUGAAUAAGGUGAA----UAGUUAAUGUGGGUUGUUUUGCCAGUUCACUUUUGACUGAGGAUCGGUUGGAUGGAGUCACCUCGACG- ....(((((..(((((.....)))))..----)))))...(((..(((......)))..))).((..((((.....))))..))....(((.....))).- ( -23.20, z-score = -0.07, R) >dp4.chrXL_group1e 6365765 94 + 12523060 ----ACUUUUCUGCUUUCGUUGUACGUA-AUGGGAUUGCGAUCAGUUCUCUUGUAGCUUCGAAAGCGAAUGGAUGCUGGGAUGCUGGGUUCCCAUGCCC-- ----.((.(((.(((((((..(((((..-..(((((((....)))))))..))).))..)))))))))).))..(((((((........))))).))..-- ( -28.00, z-score = -1.15, R) >droEre2.scaffold_4690 5900124 95 + 18748788 UUUGAGCUGCGUACUUGAGUAAGGUGAA----UAGUUAAUGUGGGUUG-UUCGCCAGCUCACUUUUGAUUGAGGAUCGGUUGGAUGGAGUCACCUCAGCG- .....((((..(((....)))((((((.----........((((((((-.....)))))))).((..((((.....))))..)).....)))))))))).- ( -27.40, z-score = -0.89, R) >droYak2.chrX 8335063 100 - 21770863 UUUGGAACGCGUACUUGAAUAAGGUGAAUAGUUAGUUAAUGUGGCUUGCUUGGCCAGGUUACUUUUGACUGGGGAUCGGUUAAAUGGAGUCAUCUCUACG- .....(((...(((((.....)))))....)))(((.(((.(((((.....))))).))))))((((((((.....))))))))(((((....)))))..- ( -24.10, z-score = -0.58, R) >droSec1.super_20 686678 96 - 1148123 UUUGAGCUGCGUACUUGAAUAAGGUGAA----UAGUUAAUGUGGGUUGUUUUGCCAGUUCACUUUUGACUGAGGAUCGGUUGGAUGGAGUCACCUCGACG- ....(((((..(((((.....)))))..----)))))...(((..(((......)))..))).((..((((.....))))..))....(((.....))).- ( -23.20, z-score = -0.07, R) >droSim1.chrX 10824889 96 - 17042790 UUUGAGCUGCGUACUUGAAUGAGGUGAA----UAGUUAAUGUGGGUUGUUUUGCCAGUUCACUUUUGACUGAGGAUCGGUUGAAUGGAGUCACCUCGACG- ((..((.......))..))((((((((.----........(((..(((......)))..))).((..((((.....))))..)).....))))))))...- ( -26.80, z-score = -1.31, R) >droMoj3.scaffold_6359 2754450 100 - 4525533 -UUAAGAUCGCCAAAUGAGAGAAAGAUUUUGAAUUUUGAUUGUUUUCAAUUUGCCACGCCUCUGCUGGCGUGUGUUUCUCUAUAUUGAUAGAUCCCUAUCU -....((((....(((((((((((.(.........((((......)))).....((((((......))))))).))))))).))))....))))....... ( -25.80, z-score = -2.95, R) >consensus UUUGAGCUGCGUACUUGAAUAAGGUGAA____UAGUUAAUGUGGGUUGUUUUGCCAGUUCACUUUUGACUGAGGAUCGGUUGGAUGGAGUCACCUCGACG_ .....................((((((.....(((((((.((((((((......))))))))..)))))))....((.((...)).)).))))))...... ( -9.14 = -8.59 + -0.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:42:18 2011