
| Sequence ID | dm3.chrX |
|---|---|
| Location | 13,821,434 – 13,821,527 |
| Length | 93 |
| Max. P | 0.500000 |

| Location | 13,821,434 – 13,821,527 |
|---|---|
| Length | 93 |
| Sequences | 12 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 83.47 |
| Shannon entropy | 0.30387 |
| G+C content | 0.46010 |
| Mean single sequence MFE | -28.14 |
| Consensus MFE | -18.78 |
| Energy contribution | -19.22 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chrX 13821434 93 - 22422827 ------GUUAACCCACGCAUGCCUGAGCAAUUUAGUGUGUUAAGGCUUUUGCACUAAAAUAAAUCUACGGCGAUUUGUUUGUUUGCACAAGCUGCCGGC------------------ ------.(((((.(((...(((....))).....))).)))))(((((.((((...(((((((((......)))))))))...)))).)))))......------------------ ( -25.70, z-score = -0.89, R) >droSec1.super_20 467529 93 - 1148123 ------GUUAACCCACGCAUGCCUGAGCAAUUUAGUGUGUUAAGGCUUUUGCACUAAAAUAAAUCUACGGCGAUUUGUUUGUUUGCACAAGCUGCCGGC------------------ ------.(((((.(((...(((....))).....))).)))))(((((.((((...(((((((((......)))))))))...)))).)))))......------------------ ( -25.70, z-score = -0.89, R) >droSim1.chrX 10635353 93 - 17042790 ------GUUAACCCACGCAUGCCUGAGCAAUUUAGUGUGUUAAGGCUUUUGCACUAAAAUAAAUCUACGGCGAUUUGUUUGUUUGCACAAGCUGCCGGC------------------ ------.(((((.(((...(((....))).....))).)))))(((((.((((...(((((((((......)))))))))...)))).)))))......------------------ ( -25.70, z-score = -0.89, R) >droYak2.chrX 8116036 93 - 21770863 ------GUUAACCCACGCAUGCCUGAGCAAUUUGGUGUGUUAAGGCUUUUGCACUAAAAUAAAUCUACGGCGAUUUGUUUGUUUGCACAAGCUGCCGGC------------------ ------......((..(((((((.((....)).)))))))...(((((.((((...(((((((((......)))))))))...)))).)))))...)).------------------ ( -28.10, z-score = -1.37, R) >droEre2.scaffold_4690 5683201 93 + 18748788 ------GUUAACCCACGCAUGCCUGAGCAAUUUAGUGUGUUAAGGCUUUUGCACUAAAAUAAAUCUACGGCGAUUUGUUUGUUUGCACAAGCUGCCGGC------------------ ------.(((((.(((...(((....))).....))).)))))(((((.((((...(((((((((......)))))))))...)))).)))))......------------------ ( -25.70, z-score = -0.89, R) >droAna3.scaffold_13047 1347882 93 - 1816235 ------GUUAACCCACGCAUGCCCGAGCAAUUUAGUGUGUUAAGGCCUUUGCACUAAAAUAAAUCUGCGGCGAUUUGUUUGUUUGCACAAGCUGCCGGC------------------ ------.(((((.(((...(((....))).....))).)))))((((((((((...(((((((((......)))))))))...)))).)))..)))...------------------ ( -24.30, z-score = -0.02, R) >droPer1.super_21 248007 87 + 1645244 ------GUUAACCCACGCAUGCCUGAGCAAUUUAGUGUGUUAAGGCUUUUGCACUAAAAUAAAUCUACAGCGAUUUGUUUGUUUGCACAAGCC------------------------ ------.(((((.(((...(((....))).....))).)))))(((((.((((...(((((((((......)))))))))...)))).)))))------------------------ ( -26.60, z-score = -3.05, R) >dp4.chrXL_group1e 6113811 87 + 12523060 ------GUUAACCCACGCAUGCCUGAGCAAUUUAGUGUGUUAAGGCUUUUGCACUAAAAUAAAUCUACAGCGAUUUGUUUGUUUGCACAAGCC------------------------ ------.(((((.(((...(((....))).....))).)))))(((((.((((...(((((((((......)))))))))...)))).)))))------------------------ ( -26.60, z-score = -3.05, R) >droWil1.scaffold_181096 8403807 116 - 12416693 GCCUAACUUGGCCCACGCAUGCCUGAGCAAUUUGGUGU-UGUAGGCUCGUGCACUAAAAUAAAUCUACGGCGAUUUGUUUGUUUGCACAAGCCGCGGCAGCAACCAGCAAGCUGGUC (((......)))....((.((((...(((((.....))-))).((((.(((((...(((((((((......)))))))))...))))).))))..)))))).(((((....))))). ( -45.10, z-score = -2.45, R) >droVir3.scaffold_12970 10059880 89 + 11907090 ------GACUACCCACGCAUGCCUGAGCAAUUUUGC---UGUGCGCUU-UGCACUAAAAUAAAUCUGCGGCGAUUUGUUUGUUUGCACAAGUCGCCGCC------------------ ------..........(((.(((..((((....)))---)..).))..-)))..............((((((((((((........)))))))))))).------------------ ( -29.00, z-score = -2.64, R) >droMoj3.scaffold_6359 1475088 92 + 4525533 ------GACUACCCACGCAUGCCUGAGCAAUUUCGC---UGUGCGCUU-UGCACUAAAAUAAAUCUGCAGCGAUUUGUUUGUUUGCACAAGUCGCCGCCGAG--------------- ------..........(((.(((..(((......))---)..).))..-)))...........((.((.(((((((((........))))))))).)).)).--------------- ( -24.00, z-score = -0.60, R) >droGri2.scaffold_15081 2706765 89 + 4274704 ------GACUGCCCACGCAUGCCAGAGCAAUUUCGC---AGUGCGCUU-UGCACUAAAAUAAAUCUGCGGCGAUUUGUUUGUUUGCACAAGUCGCCGCC------------------ ------...(((...(((((((.(((....))).))---).))))...-.))).............((((((((((((........)))))))))))).------------------ ( -31.20, z-score = -2.33, R) >consensus ______GUUAACCCACGCAUGCCUGAGCAAUUUAGUGUGUUAAGGCUUUUGCACUAAAAUAAAUCUACGGCGAUUUGUUUGUUUGCACAAGCCGCCGGC__________________ ................((.(((....)))..............(((((.((((...(((((((((......)))))))))...)))).)))))))...................... (-18.78 = -19.22 + 0.44)


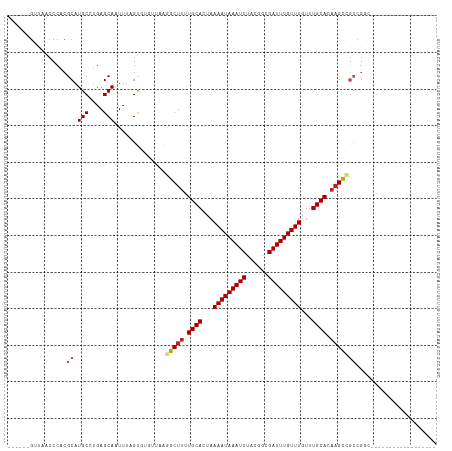
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:41:34 2011